AI Unleashes
the Power of Data
Unlock insights from your sensor data with AI, Generative AI, and Machine Learning to
get accurate, actionable responses to user prompts to make smarter decisions.
Skin Cancer Detection
Computer Vision – Health History

Gen AI Travel
Full Stack & Grok Integration

Time Series ML & Gen AI
Cognitive & Musculoskeletal

Gen AI Biometric Eval
Improved Accuracy Blood Pressure
Grok Generative AI Prompt / Fine Tuning

Travel Recommendations
Travel with Experience-First AI
Hybrid: GraphSage GNN / LLM

Transformers
Large Language Model
HuggingFace – Full Model

^ Human Productivity
n8n Customized AI Agents
Coming Soon

Personal Data Sovereignty
Retrieval-Augmented Generation
Private Brain Semantic Search

AI Coding Workflow
AI – Assisted and Augmented
Sovereign Bible + Notes RAG
Skin Cancer Diagnosis – AI Computer Vision
Goal: Develop a rapid home test to properly classify benign vs. malignant skin lesions and convince my father to get a suspect skin lesion inspected by a Dermatologist
- This method utilizes machine learning to classify images (benign or malignant) and medical history to enhance classification. It is hypothesized that history may be more useful than simply optimizing the accuracy of the model and will save time in development
- Additional methods using skin masking (image segmentation using DeepLabV3) and an ensemble approach were utilized for improving accuracy
- Input Requirements: A) Method to capture a picture (i.e. smart phone) and access to the internet and B) short survey regarding history of skin lesion
- Output: Malignant vs. Benign
Methods:
- Random images from the ISIC 2020 Dataset (The International Skin Imaging Collaboration) with varied participants (sex, age, nationality – white skin tone)
- Training set of 457 benign and 457 malignant (total: 914) images and test set of 125 benign and 125 malignant (total: 250) images
- Images in JPG format were processed from various landscape pixel widths and heights to square 299 X 299-pixel RGB images with one model using 254 pixels square and another using 224 pixels square.
- The most relevant portion of the image was maintained during cropping
- Hair was not removed and in some cases scale measurement marks were left
- Various supervised, convolutional neural network (CNN) computer vision models were evaluated: Custom Encoder / Decoder Model, Inception V3, Xception, ResNet18, ResNet50, ResNet152, VGG16, VGG19, EfficientNet_B2, and DenseNet121
- Hyperparameters were tuned, data normalized, and training data was augmented (randomly horizontally flipped, partially rotated, size / contrast changed, etc.)
A short survey was given to learn the history of the skin legion (case studies)
- How long was the skin legion present?
< 3 months, < 6 months, < 1 year, < 5 years, lifetime - Is skin legion greater than size of a single standard No 2 pencil end eraser (6 mm)?
Yes or no - Has the skin legion increased in size over time? Yes or No
- How much as the skin legion increased in size?
< Doubled in size, Doubled in size, > Doubled in size - How long ago did it start growing? 3 months, 6 months, 1 year, 5 years, slowly over lifetime
- Does it bleed on it’s own (without picking, squeezing)? Yes or No
- Has it significantly changed in appearance? Yes or No
- Have you had a previous malignant skin lesion(s) (skin cancer) removed? Yes or no
- Does lesion itch, burn or cause pain? Yes or No
This information is not available from ISIC database and is only available for case studies.
First Step in AI is Understanding the Data
Morphological Characteristics of a Malignant Skin Lesion

Asymmetry

Border Irregularity

Color / Raised

> 6 mm Diameter
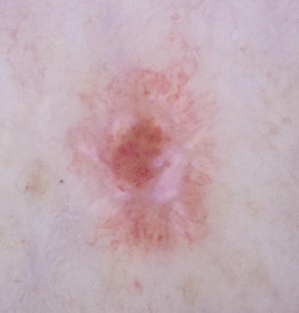
Ring or Halo of Redness
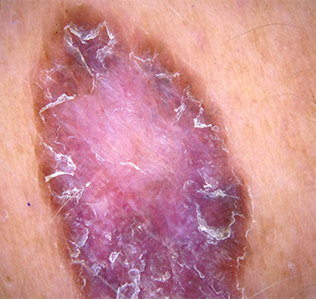
Scales

Ulcerations / Crust

Evolution Over Time
Examples of Malignant Skin Lesions
Source: National Cancer Institute
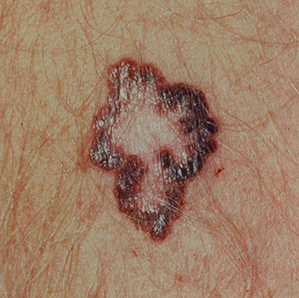
Melanoma

Melanoma

Asymmetrical Melanoma
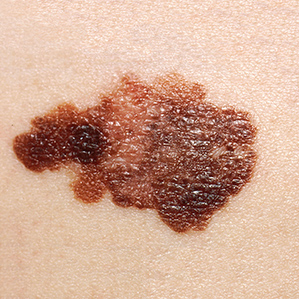
Melanoma
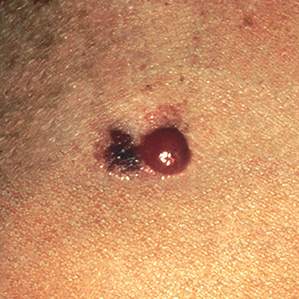
Melanoma Advanced Stage

Melanoma

Basal Cell Carcinoma Ulcerated
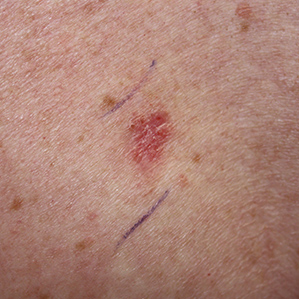
Basal Cell Carcinoma Superficial

Basal Cell Carcinoma

Basal Cell Carcinoma
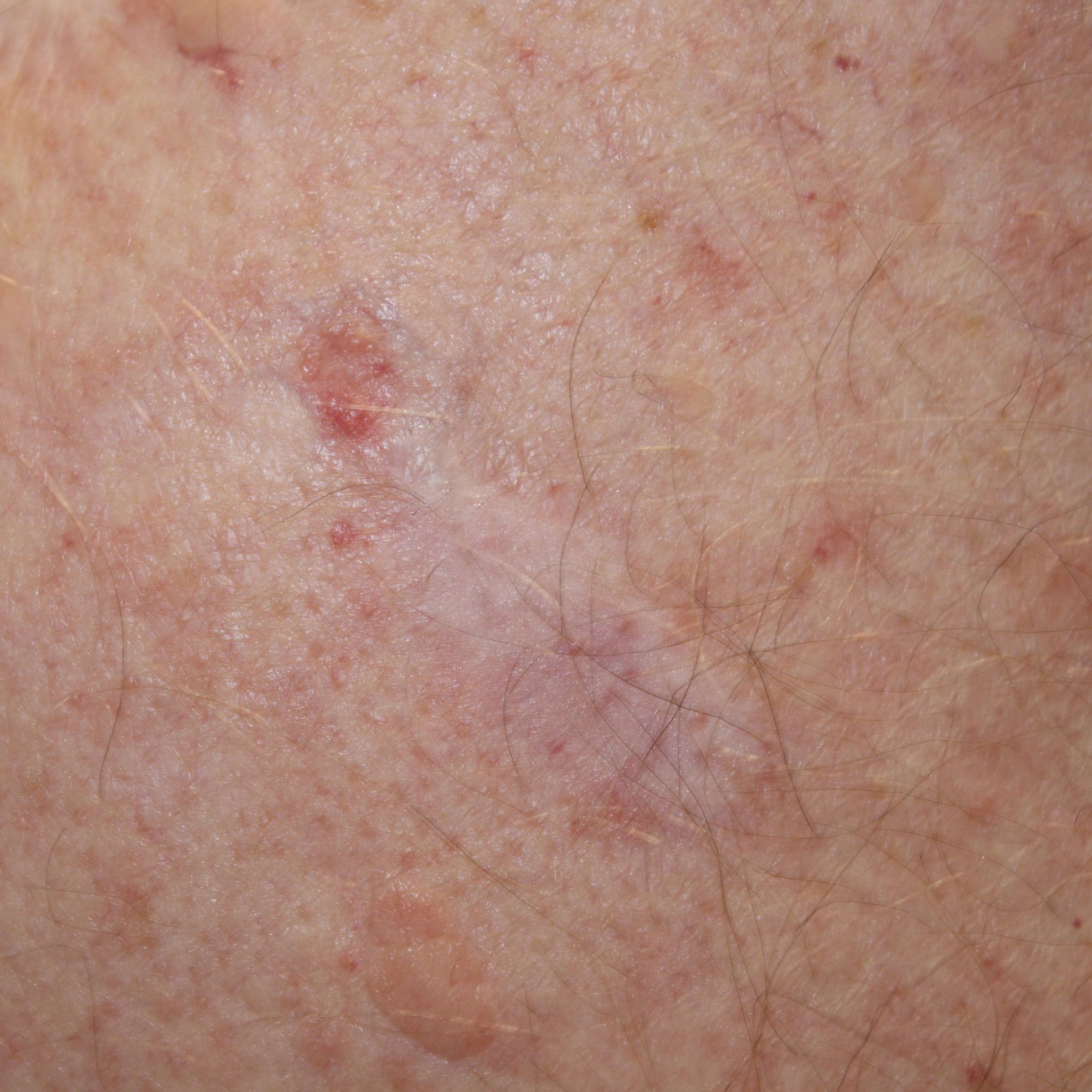
Basal Cell Carcinoma
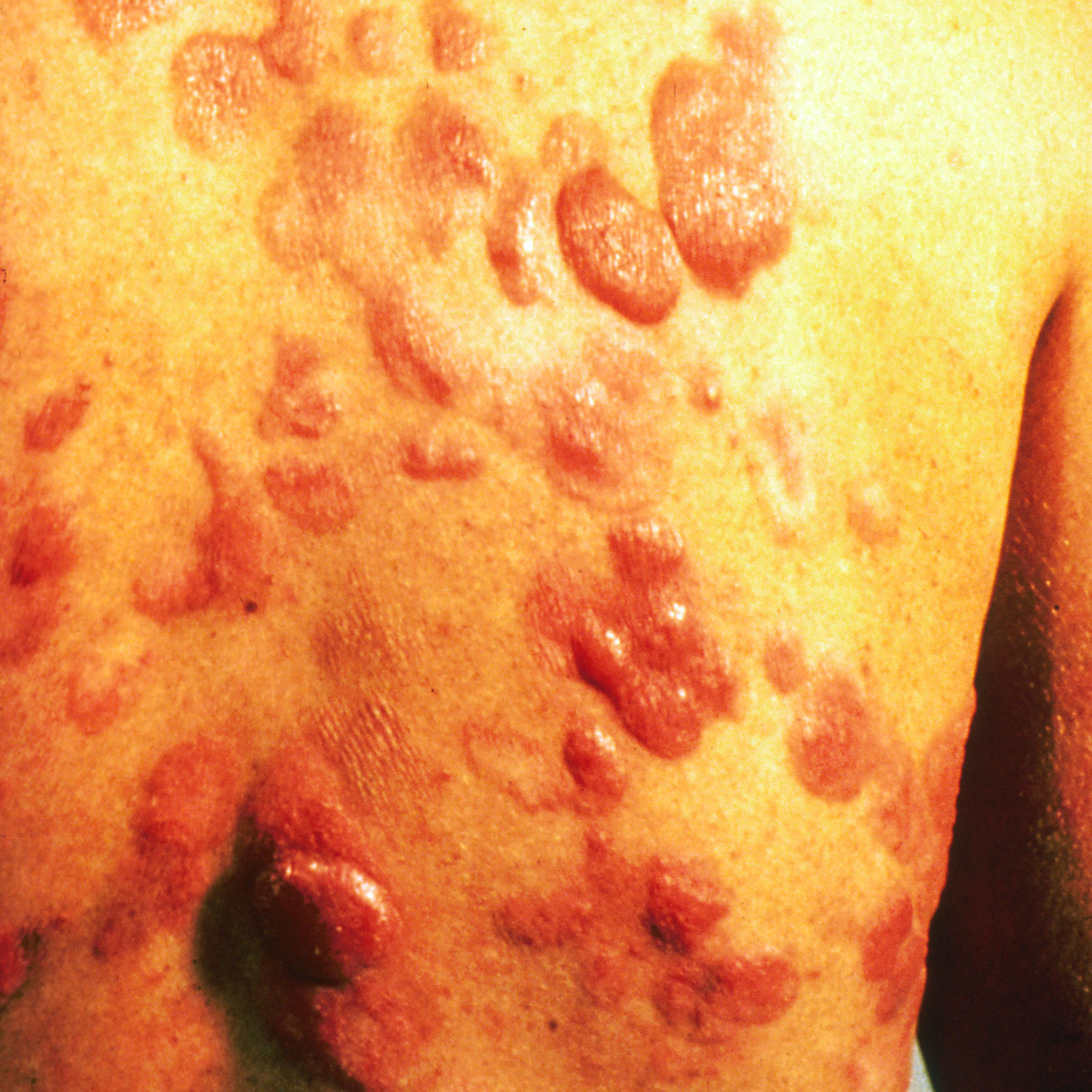
Kaposi’s Sarcoma
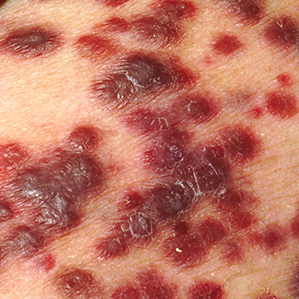
Kaposi’s Sarcoma
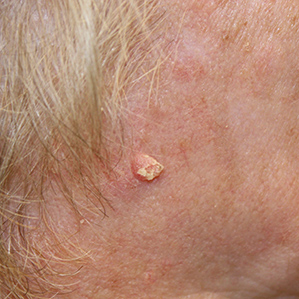
Squamous Cell Carcinoma

Squamous Cell Carcinoma
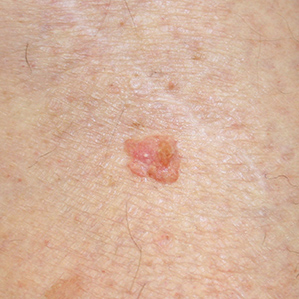
Squamous Cell Carcinoma
Examples of Masked Images (Image Segmentation with DeepLabV3)
Using binary segmentation image processing, the image was divided into two distinct regions: 1) the skin lesion to be classified as maglinant or benign and 2) the surrounding healthy skin. The surrouding healthy skin was then masked so only the skin lesion was visible. There are a number of methods to accomplish this including Meta’s Segment Anything Model (DICE of 0.93), nnU-Net (DICE of 0.96), and TransUNet (DICE of 0.95) but each of these would have taken many hours to fine tune and required significant GPU resources. BASNet or Boundary-Aware Segmentation Network was also considered (2 – 4 hours of training, DICE of 0.98) but I preferred DeepLabV3 which was readily available in PyTorch and could be trained in minutes without another software download. DeepLabV3 is an atrous spatial pyramid pooling (ASPP) model, designed for semantic segmentation and is widely used in medical imaging. ASPP applies several atrous convolutions in parallel, each with a different dilation rate (e.g., 6, 12, 18), to capture features at multiple scales: 1) small features with low dilation rates (e.g., 6) focus on fine details, like lesion edges or small lesions and 2) large features with high dilation rates (e.g., 18) to capture broader context, like the overall shape or surrounding skin texture. The DeepLabV3 model used ground truth masks (DICE 1.0) from the ISIC 2018 dataset for training. The Dice coefficient is a statistical metric used to quantify the accuracy of image segmentation.

Benign
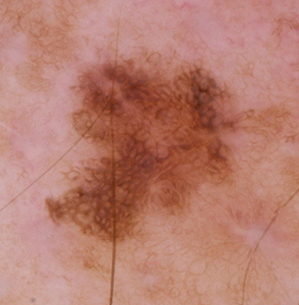
Benign

Malignant
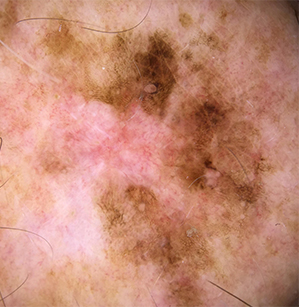
Malignant

Benign Masked
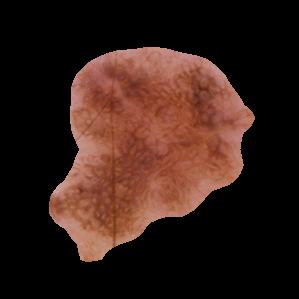
Benign Masked
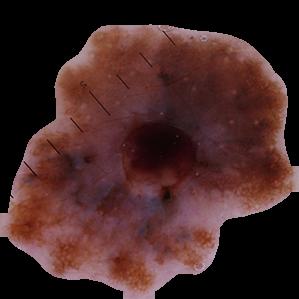
Malignant Masked

Malignant Masked
Example Python Code for EfficientNet_B2 in Pytorch
This is an example of code to train a convolutional neural network (EfficientNet_B2) model. Several other models were considered including ensembles. This provides the structure for the code without being verbose with all the code that was experimented with or used. The choice of models was limited to lower complexity models such as EfficientNet_B2 (9.1 million parameters, image size 260 x 260 pixels) vs EfficientNet_B6 (43 million parameters, image sizes up to 456 x 456 pixels) for run time efficiency.
import os
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from PIL import Image
import torchvision.transforms as transforms
from torchvision.models import efficientnet_b2
from torchvision.models import densenet121
import numpy as np
import random
random.seed(42)
np.random.seed(42)
torch.manual_seed(42)
torch.cuda.manual_seed_all(42)
# Focal Loss
class FocalLoss(nn.Module):
def __init__(self, gamma=2.0, weight=None):
super().__init__()
self.gamma = gamma
self.weight = weight
def forward(self, inputs, targets):
ce_loss = nn.functional.cross_entropy(inputs, targets, weight=self.weight, reduction='none')
pt = torch.exp(-ce_loss)
focal_loss = ((1 - pt) ** self.gamma * ce_loss).mean()
return focal_loss
# Custom Dataset
class SkinLesionDataset(Dataset):
def __init__(self, benign_dir, malignant_dir, transform=None):
self.benign_dir = benign_dir
self.malignant_dir = malignant_dir
self.transform = transform
self.images = []
self.labels = []
# Load benign images (label 0)
for img_name in os.listdir(benign_dir):
if img_name.endswith(".jpg"):
self.images.append(os.path.join(benign_dir, img_name))
self.labels.append(0)
# Load malignant images (label 1)
for img_name in os.listdir(malignant_dir):
if img_name.endswith(".jpg"):
self.images.append(os.path.join(malignant_dir, img_name))
self.labels.append(1)
def __len__(self):
return len(self.images)
def __getitem__(self, idx):
img_path = self.images[idx]
label = self.labels[idx]
image = Image.open(img_path).convert("RGB")
if self.transform:
image = self.transform(image)
return image, label
# Transforms for 299x299 images
train_transform = transforms.Compose([
transforms.Resize(256),
transforms.RandomResizedCrop(224, scale=(0.9, 1.0)),
transforms.RandomRotation(15),
transforms.RandomHorizontalFlip(p=0.5),
transforms.RandomVerticalFlip(p=0.5),
transforms.ColorJitter(brightness=0.1, contrast=0.1, saturation=0.1, hue=0.05),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
test_transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
# Datasets
train_dataset = SkinLesionDataset(
benign_dir="E:/skin/Train_With_Mask/benign",
malignant_dir="E:/skin/Train_With_Mask/malignant",
transform=train_transform
)
test_dataset = SkinLesionDataset(
benign_dir="E:/skin/Test_With_Mask/benign",
malignant_dir="E:/skin/Test_With_Mask/malignant",
transform=test_transform
)
# DataLoaders
train_loader = DataLoader(train_dataset, batch_size=64, shuffle=True)
test_loader = DataLoader(test_dataset, batch_size=64, shuffle=False)
# Model: EfficientNet-B2
model = efficientnet_b2(weights='IMAGENET1K_V1')
model.classifier = nn.Sequential(
nn.Dropout(p=0.5, inplace=True), #0.3 - (0.5 85.6%)
nn.Linear(model.classifier[1].in_features, 2) # Binary classification (benign/malignant)
).cuda()
# Unfreeze the last two MBConv blocks (stages 6 and 7)
for param in model.features[6:].parameters():
param.requires_grad = True
model = model.cuda()
# Optimizer: AdamW
optimizer = optim.AdamW(
model.parameters(),
lr=2e-4,
weight_decay=1e-3,
betas=(0.9, 0.999)
)
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, mode='max', factor=0.7, patience=3, verbose=True) #"min"
criterion = nn.CrossEntropyLoss(label_smoothing=0.1)
# Training loop
num_epochs = 6
for epoch in range(num_epochs):
model.train()
train_loss = 0.0
train_correct = 0
train_total = 0
for images, labels in train_loader:
images, labels = images.cuda(), labels.cuda()
optimizer.zero_grad()
outputs = model(images)
loss = criterion(outputs, labels)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
train_loss += loss.item()
_, predicted = torch.max(outputs, 1)
train_total += labels.size(0)
train_correct += (predicted == labels).sum().item()
train_loss /= len(train_loader)
train_acc = train_correct / train_total
# Validation
model.eval()
my_list = []
val_loss = 0.0
val_correct = 0
val_total = 0
with torch.no_grad():
#for images, labels in test_loader:
for batch_idx, (images, labels) in enumerate(test_loader):
images, labels = images.cuda(), labels.cuda()
outputs = model(images)
loss = criterion(outputs, labels)
val_loss += loss.item()
_, predicted = torch.max(outputs, 1)
val_total += labels.size(0)
val_correct += (predicted == labels).sum().item()
# Identify misclassified images
mask = predicted != labels # Creates a boolean tensor where True indicates a misclassification
for idx in range(len(mask)):
if mask[idx]:
# Calculate the global index in the dataset
global_idx = batch_idx * test_loader.batch_size + idx
# Get the filename from the dataset's images attribute
filename = test_loader.dataset.images[global_idx] # Use images attribute
my_list.append(filename)
val_loss /= len(test_loader)
val_acc = val_correct / val_total
print(f"Epoch {epoch+1}, Train Loss: {train_loss:.4f}, Train Acc: {train_acc:.4f}, "
f"Val Loss: {val_loss:.4f}, Val Acc: {val_acc:.4f}")
scheduler.step(val_loss)
# Print the filenames of misclassified images
print("Misclassified image filenames:")
for filename in my_list:
print(os.path.basename(filename))
print(f"Number of misclassified images: {len(my_list)}")
# Save model
torch.save(model.state_dict(), "E:/skin/models/efficientnet_b2_finetuned.pth")
print("Model saved to E:/skin/models/efficientnet_b2_finetuned.pth")
EfficientNet_B2: Loss – Accuracy Graphs

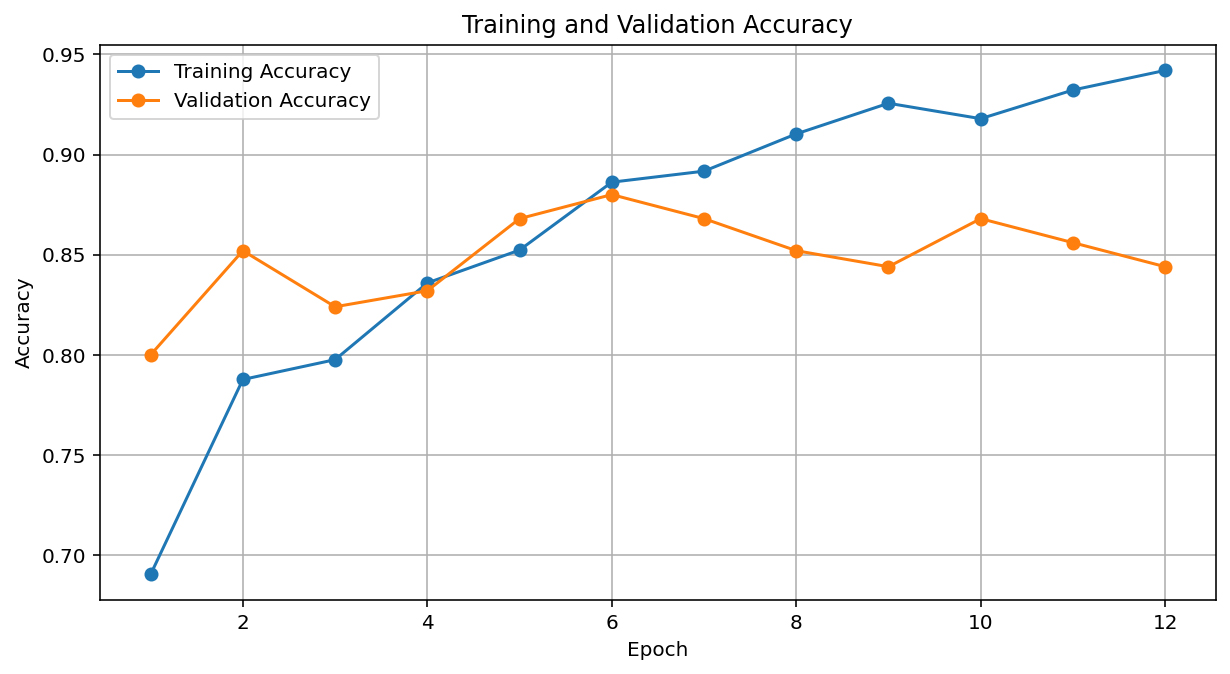
Computer Vision Results
- It is noted that the vast majority of malignant skin lesion images were melanoma.
- Three individual models were chosen for their training and evaluation speed and accuracy: 1) ResNet18 (86.8% accuracy), 2) EfficientNet_B2 (88.0% accuracy) and 3) DenseNet121 (87.6% accuracy). EfficientNet_B2 and DenseNet121 models used masked images and ResNet18 used the full image with no masking.
- An ensemble of DenseNet121, ResNet18, and EfficientNet_B2 (90.0% accuracy) was also evaluated. The ensemble made predictions with the highest accuracy EfficientNet_B2 model and only changed it’s result if ResNet18 and DenseNet121 agreed on an alternative prediction.
- Sample size was limited due to computing resources and hence accuracy was not optimized beyond this point. Various other ensemble approaches (softmax averaging, majority voting, weighted voting, etc.) were considered but the one highlighted above was the most accurate.
- The malignant accuracy was better than the benign accuracy for each of the models. For example, with EfficientNet_B2 benign accuracy was 83.2%
and malignant accuracy was 92.8%. - False positives from analysis of images alone: 7.2% (cautious approach, high likelihood of biopsy). See sampling (not all) below.
- False negatives from analysis of images alone: 3.2% (risk of not being treated resulting in high potential for adverse outcome). See sampling (not all) below.
- Most false positives very difficult to determine from image alone hence examining and including health history has merit.
- 87.5% of false negatives I would have gotten biopsied from image alone hence accuracy improvement in model is warranted with more training images.

False Positive
Benign

False Positive
Benign

False Positive
Benign

False Positive
Benign
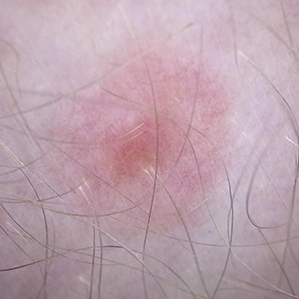
False Positive
Benign
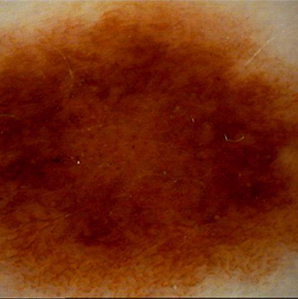
False Positive
Benign

False Positive
Benign
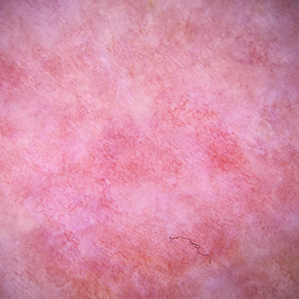
False Positive
Benign

False Positive
Benign

False Positive
Benign

False Positive
Benign

False Positive
Benign
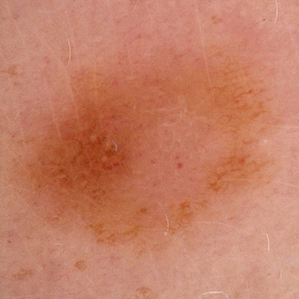
False Negative
Malignant

False Negative
Malignant
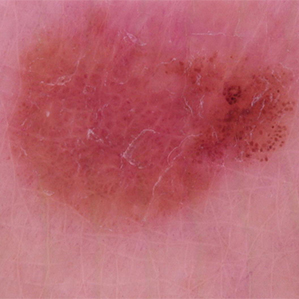
False Negative
Malignant
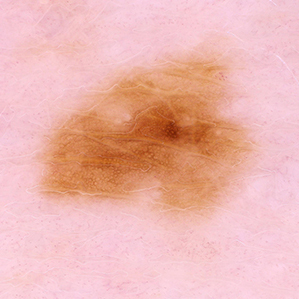
False Negative
Malignant
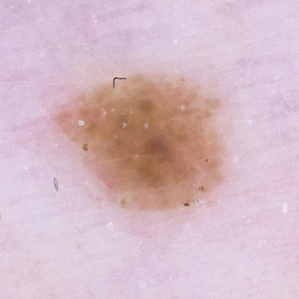
False Negative
Malignant

False Negative
Malignant
Case Studies

Mole from Birth
Model: Benign
Health History: Benign

Birthmark from Birth
Model: Malignant
Health History: Benign
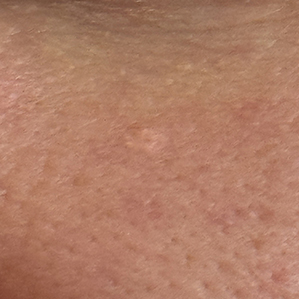
New Growth Face
Model: Malignant
H. History: Malignant

Large Mole Many Years
Model: Benign
Health History: Benign

New Large Lesion
Model: Malignant
H. History: Malignant
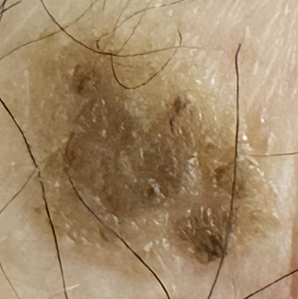
New Large Lesion
Model: Malignant
H. History: Malignant
The AI model was correct for 3 / 6 case studies and possibly correct but unconfirmed for 1/6. It made two mistakes on an extremely challenging birthmark and a new growth on my face. The new growth on my face was Sebaceous hyperplasia or an oil-producing sebaceous gland that became enlarged / clogged (layman’s terms) and resembled Basal Cell Carcinoma which I had previously at a younger age. The models had a higher tendency for false positives than false negatives which was good. EfficientNet_B2 accurately diagnosed this lesion as benign but since DenseNet121 and ResNet18 both diagnosed this as malignant, the EfficientNet results was overturned. Hence, it was prudent that I got it checked. The birth mark resembled melanoma but health history proved to be very important in this case. Given the subjects age, no history of skin cancer, and being present since birth, this legion was not suspect and this was confirmed by a dermatologist. The first new large lesion was a confirmed (biopsy) cancerous lesion on the arm. The last case study is a new enlarged lesion on my dad’s chest. Despite my efforts with this analysis, the malignant diagnosis of the model, and his history of skin cancer, he will not get it checked. He is at end of life and doesn’t have an interest so the case is not confirmed by biopsy.
Overall, health history definitely helped in diagnosis but also contributed to a false positive. The case studies also confirmed what was already known in that a much larger sample size is needed to improve the models from 90 to 95-96% accuracy (best in the literature). It would be very interesting to test health history in conjunction with model performance on 5000 samples. I would also like to test more complex diagnosis and masking models with more computing resources.
Generative AI Travel
This generative AI case study integrated Grok xAI into an AWS environment and was deployed with an Amplify mobile app with ReactJS so it works on any platform. The API for Grok integration was built in the backend with Lambda, API Gateway, and Identity and Access Management. The data was stored in a S3 data lake (images) and a Dynamodb data base. The prototype was built for demonstration to Spirit Airlines prior to their bankruptcy. Their intention was to revolutionize their frequent flier program and AI was piece they were exploring.
Grok integration provided access to a capable large language model for user prompt engagement (Grok 1.0 used) and it’s response was enriched with images, travel features, menus, and more to enable quick decision making.
The video shows a brief demonstration of user login, the mobile app (first showing some of the health app I created), and how the travel planning process could be enhanced with generative AI. Ultimately this can be improved by aiding the user in developing a travel itinerary and expanding into many elements of the travel planning process. Spirit specifically requested a demonstration for hotel planning.
Current travel sites leave so much to be desired with planning taking hours / days and this process can be significantly enhanced with AI as a helper in the background without over dominating the experience. Too many times AI completely takes over leaving the user with a frustrated experience. Allowing the user to maintain control is extremely important.
Time Series Machine Learning and Generative AI
Problem Statement: In medicine, there is no clinical means to accurately diagnose musculoskeletal conditions such as a rotator cuff tear without advanced radiology even by a highly trained orthopedic surgeon. Furthermore, there is no uniform testing program that exists for traumatic brain injury (TBI) / cognitive conditions that includes both a means of immediate assessment on the field and comprehensive clinical management using a validated instrument.
Solution: To address this, machine learning was applied to motion signatures for the purpose of medical triage, diagnosis, and monitoring to facilitate expedient, cost efficient, and effective patient care and health management for musculoskeletal and cognitive conditions. The motion signatures are generated from data collected from inertial sensors monitoring dynamic motion of the human body. Furthermore, generative AI was introduced in combination with machine learning classification and regression of motion signatures for treatment planning and injury prediction. The generative AI is a fine-tuned LLM, pretrained in a broad corpus of medical knowledge and further refined using domain-specific datasets including AI-driven motion signature profiling, biometric profiles, and health records. This multimodal approach provides a novel data-driven framework for comprehensive patient assessment and treatment planning using diverse inputs through its generative architecture to produce actionable outputs. For example during triage, the LLM evaluates the combined data with a high anomaly score, elevated pain, and a history of prior injury to assign a priority level (e.g., urgent, routine), guiding clinical resource allocation.
To validate this AI-based system, a 35-person clinical study was conducted for rotator cuff injury with 96% accuracy in classification of rotator
cuff tear vs. inflammation without advanced radiology although MRIs were used to determine proper labeling of the training and test data. In addition, another groundbreaking study for diagnosing TBI and degenerative cognitive conditions was conducted leveraging cognitive impairment data (sober vs intoxicated), with an initial model accuracy of 80% gated by limited sample size and an interesting fact. Careful review of the data showed men lost more motor control than women with intoxication. Hence the machine learning properly classified all sober subjects and men intoxicated, but not women intoxicated. The specifics of the algorithms for this patent pending AI invention, owned by David, cannot be discussed in detail but the data and general approach will be shared as presented at national conferences.
Test Methods Musculoskeletal Injury
- Random subjects (men and women) with varied ages participated. This resulted in a sample size of 35 divided between train and test groups for the machine learning algorithm. There were 18 normal (inflammation only) and 17 abnormal conditions (rotator cuff tear).
- Two tests were conducted: Forward extension and external rotation.
- Forward extension: Arm at side and elevated in the Scapular plane (45 degrees between front and side position, fully extended) 180 degrees overhead and back down at side.
- External rotation: Arm positioned in an “L” orientation with the elbow touching the hip and then rotated front to back and back to front.
- The tests were conducted with no weight and weight in the hand.
- Sensor was located in the sulcus of the inner biceps and secured with a strap.
Test Methods Cognitive Impairment
- Random subjects (60% men; 40% women) with varied ages participated. This resulted in a sample size of 20 divided between train and test groups for the machine learning algorithm.
- Participants were asked to perform two tactile edge orientation processing movements, sober and under the influence of alcohol. This included a lower extremity leg movement (LELM) test of the heel sliding along the tibia of the opposite leg from bottom of shin to knee and back (in seated position) and the “what time is it?” test with movement of arm from side, reading time of wristwatch, and returning to the side.
- Cognitive conditions were simulated with BAC (blood alcohol content) levels of 0.00 (sober), 0.02, 0.04, 0.06, 0.08, and 0.09%. The BAC was measured using a breathalyzer. Each motion test was repeated three times.
- Sensors were located either on the wrist (next to watch) for the “what time is it?” test or on the inside ankle for the LELM test (strap made transparent so you can see sensor).
Sensor Locations
External Rotation / Forward Extension

What Time is It?

Lower Extremity Leg Movement

Data from Musculoskeletal Evaluations
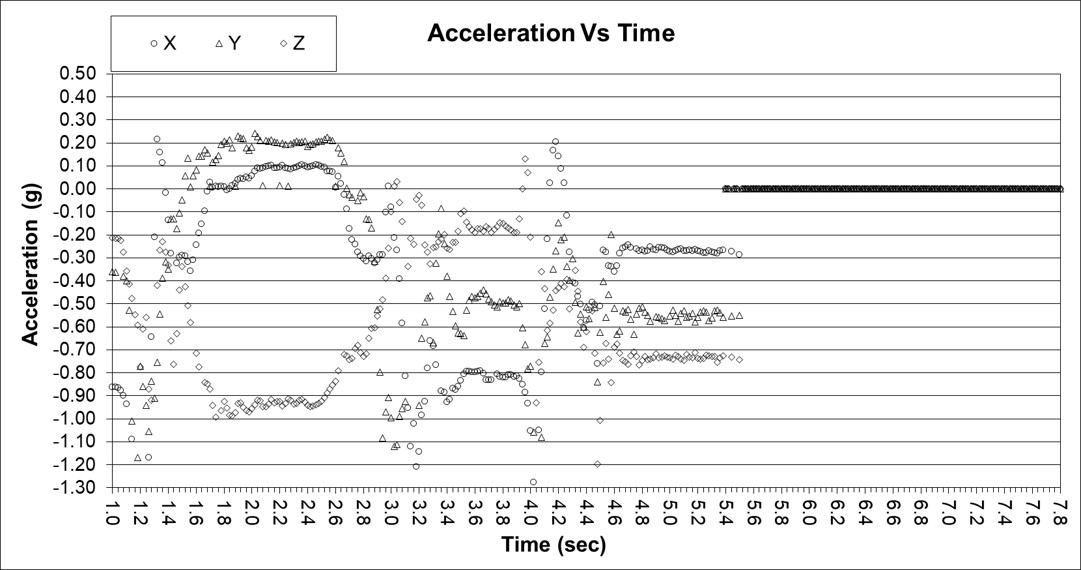
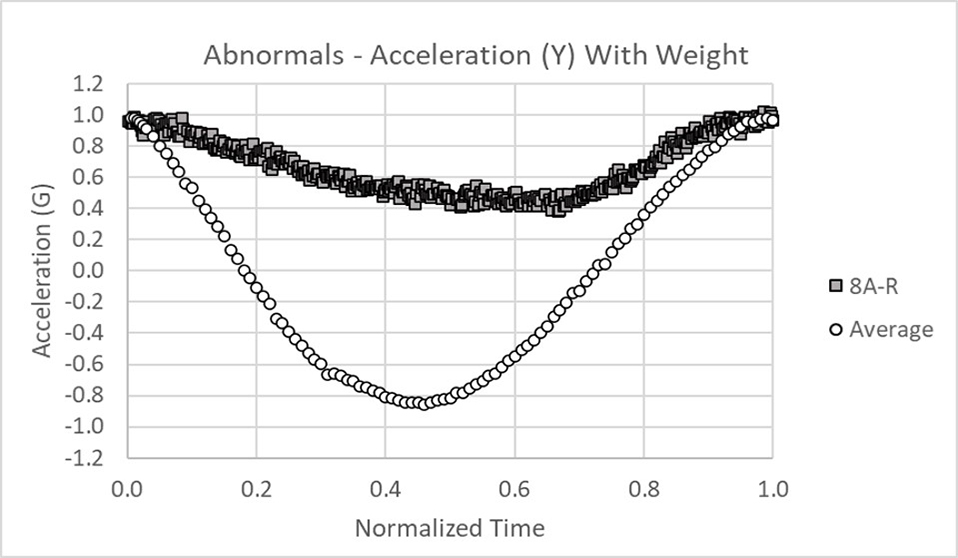
Data was collected in the x, y, and z axes for acceleration and rotation of the arm in forward extension. The data below shows 18 normals tightly banded around the average and +/- 3 sigma limits (dark points). Only the acceleration data in the x and y axes is shown for brevity.


This is an example of abnormal acceleration data for the x and y axes. Clearly the data falls outside of the +/- 3 sigma limits.


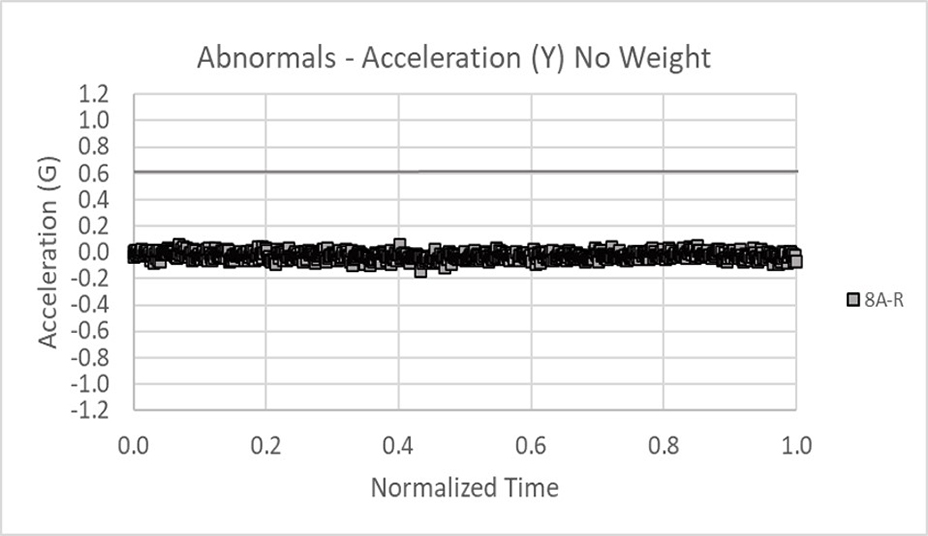
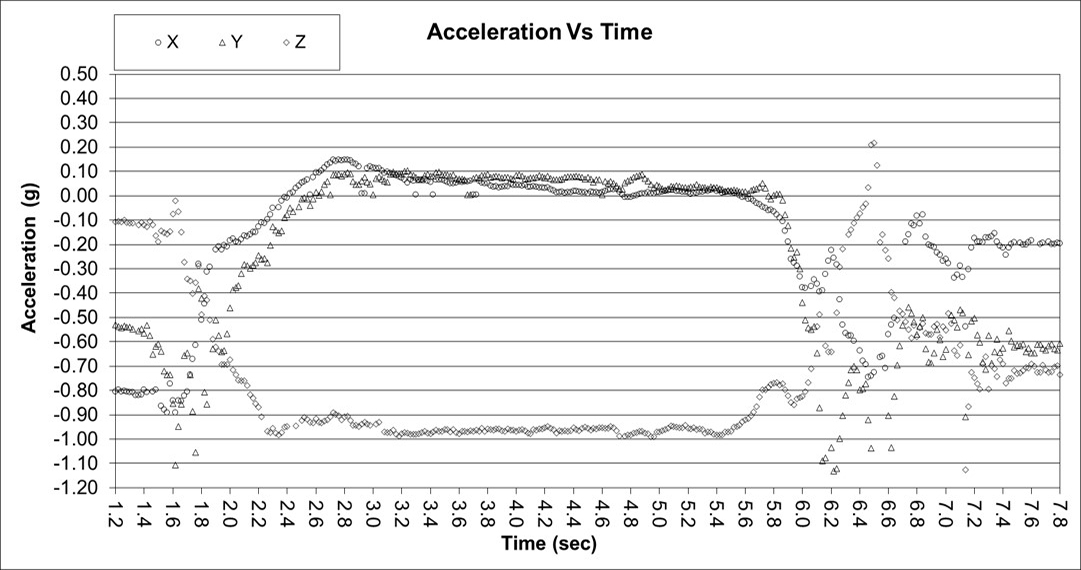
Data was collected in the x, y, and z axes for acceleration and rotation of the arm in external rotation. The data below shows 18 normals tightly banded within the dark lines (not +/- 3 sigma limits but critical thresholds). Only the acceleration data in the x and y axes is shown for brevity.
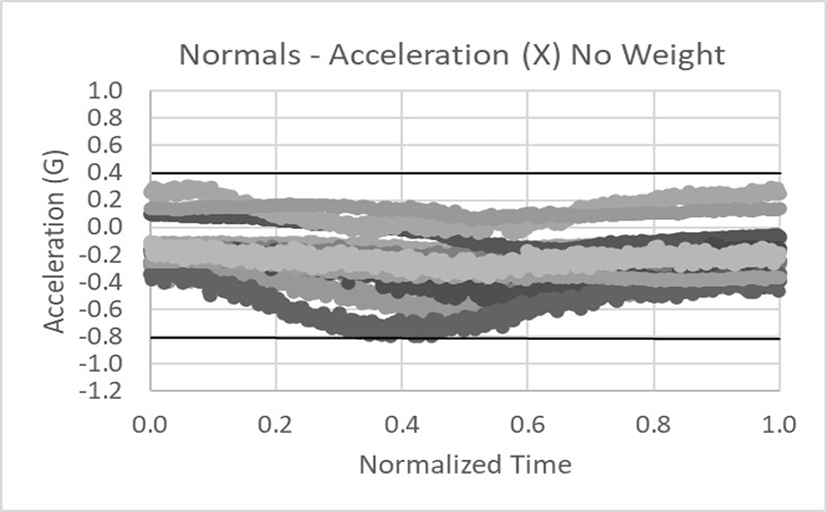
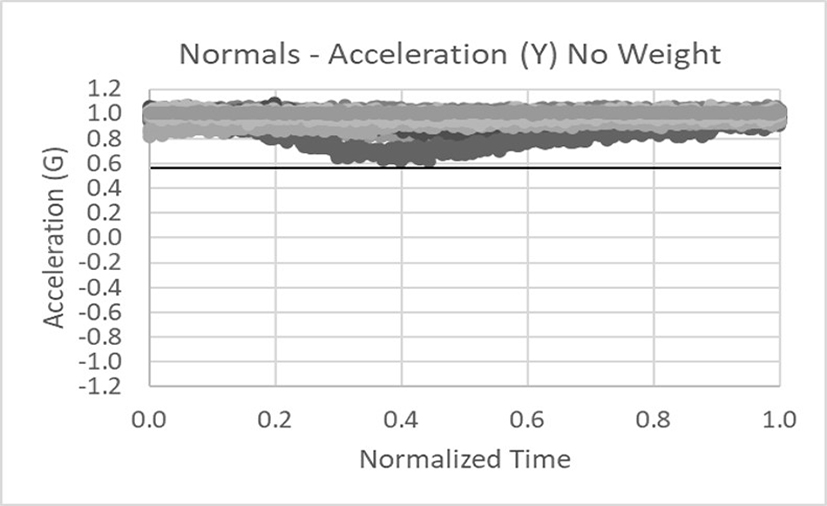
This is an example of abnormal acceleration data for the x and y axes. Clearly the data falls outside of the threshold limits.

Data from Cognitive Evaluations
Data was collected in the x, y, and z axes for acceleration and rotation of the arm during the “What time is it?” test. The data below shows a clear difference between sober and intoxicated. Only a portion of the acceleration data shown.
Sober BAC 0.00%
Intoxicated BAC 0.09%
Data was collected in the x, y, and z axes for acceleration and rotation of the leg during the lower extremity leg movement test. The data below shows a clear difference between sober and intoxicated. Only a portion of the acceleration data shown.
Sober BAC 0.00%
Intoxicated BAC 0.09%

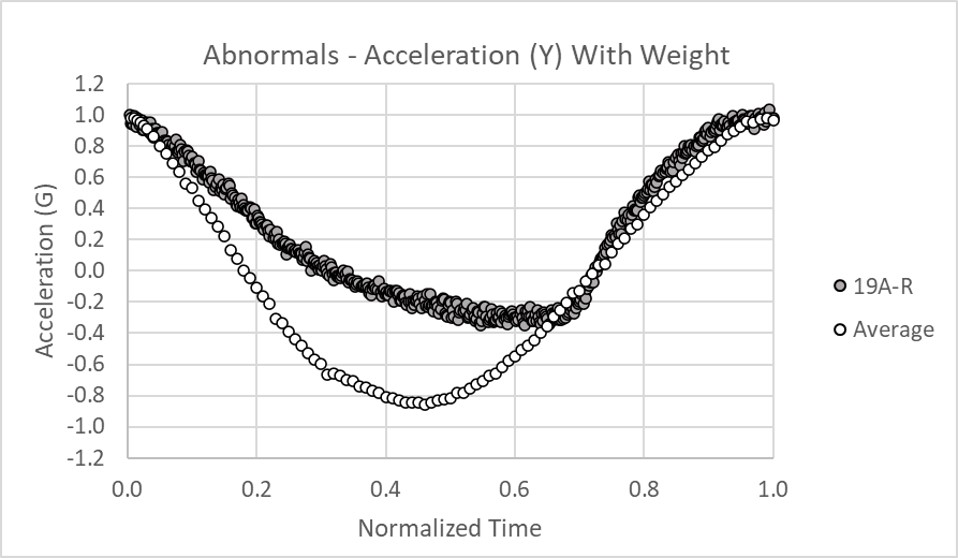
Further Exploration of the Data
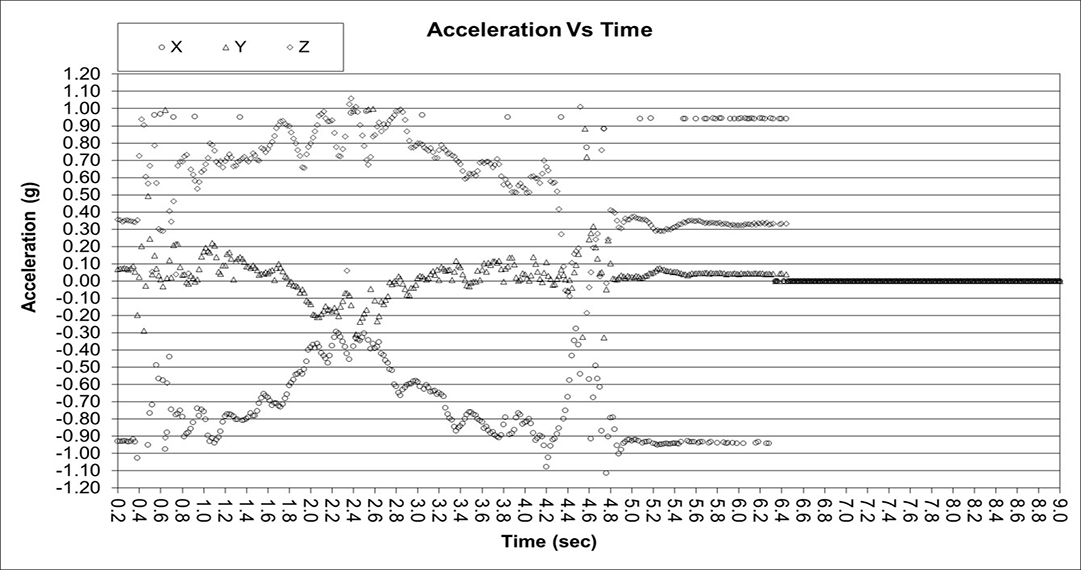
It may appear that normal (inflammation only) and abnormal (tears) rotator cuffs and sober and intoxicated are easy to distinguish with even basic statistics. However, the following data will show that a robust solution is far from trivial. The ultimate goal is to distinguish between minor partial tears (25% or less) that may not need surgery and partial tears > 50% or full tears that may need surgery. The first two graphs show clear delineation between a partial and full tear as expected but in the third graph, the full tear appears normal. However, in external rotation there is a distinguishable difference or feature between partial and full tears that traditional statistics cannot properly classify where AI excels with methods such as random shapelet transform with random forest classifiers and convolutional neural networks. If the pending patent is accepted, this subtle difference can be shared.
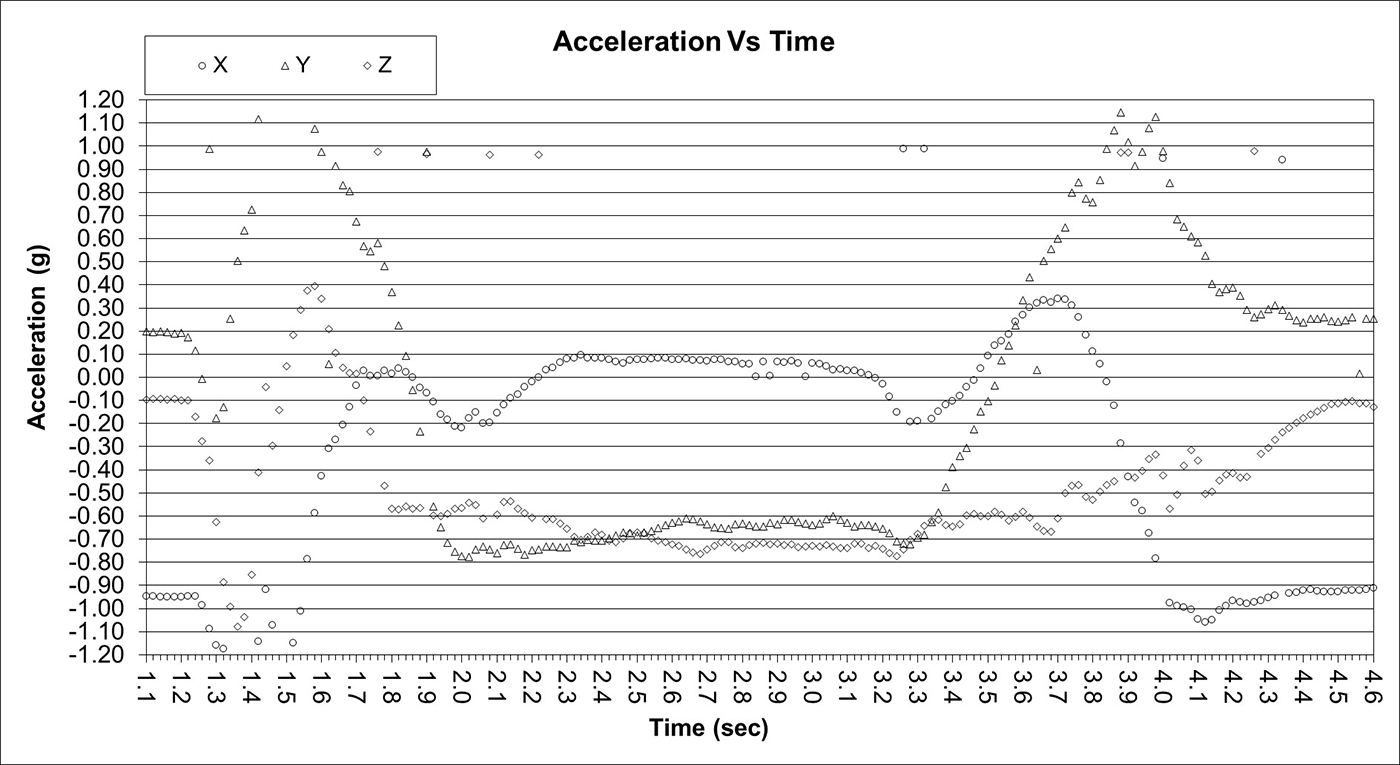
Partial Tear
Full Tear
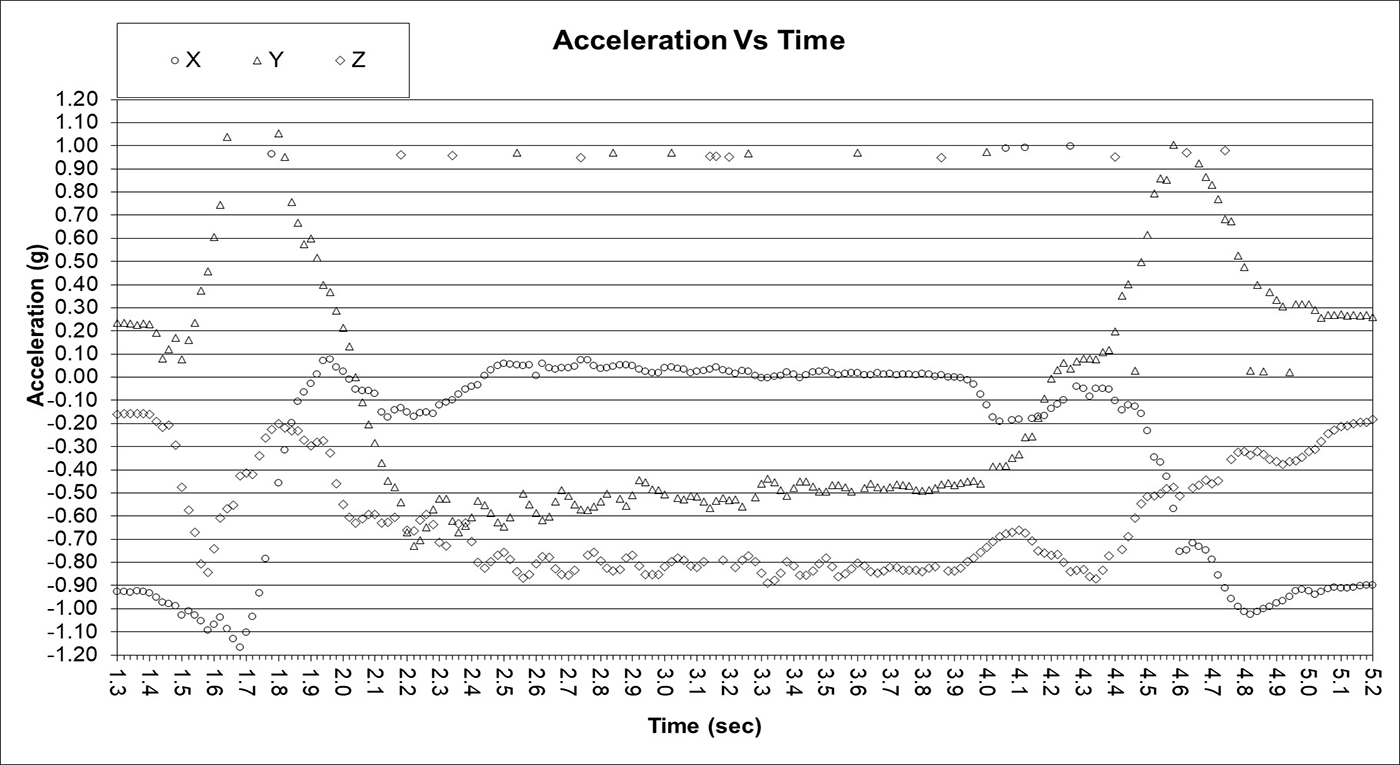
Full Tear Indistinguishable from Normal

Cognitive data of sober and intoxication showed a similar trend. In some cases, sober versus intoxicated was extremely obvious as shown above. In other cases, only subtle features were distinguishable via time series classification using random shaplet transform with random forest.
Sober BAC 0.00%
Sober BAC 0.00%
Sober BAC 0.08%

Sober BAC 0.00%

Intoxicated BAC 0.00%

Sober BAC 0.08%

Random Shapelet Transform (RST) is a technique used in time series classification that leverages subsequences called shapelets to extract meaningful, discriminative features from time series data. Shapelets are a short subsequence (a contiguous segment) of a time series data that represents a distinctive pattern or feature relevant to the classification task. It’s particularly effective for identifying local patterns or shapes within time series that are discriminative for classification tasks, such as distinguishing between normal and abnormal motion signatures. The motion signature is randomly sampled to make it faster and more scalable instead of exhaustively searching all possible subsequences (which is computationally expensive). RST then picks the best shapelets based on their ability to separate the classes (normal vs. abnormal).
For example with forward extension for rotator cuff evaluation in the y-axis, a shapelet capturing a cosine-like dip may be prevalent in normal signatures, while a shapelet capturing a central leveling may be indicative of abnormal signatures. These selected shapelets are not averages of the respective classes but are individual subsequences chosen for their discriminative power.
For each selected shapelet, the method computes a distance metric, such as the Euclidean distance, between the shapelet and every subsequence of the same length within Tᵢ (representation of time series). This is performed using a sliding window approach, where the shapelet is compared to all possible contiguous segments of Tᵢ, regardless of position. The minimum distance across all subsequences is retained for each shapelet, representing the closest match between the shapelet’s pattern and any portion of Tᵢ. This process generates a feature vector (transform) for Tᵢ, where each element corresponds to the minimum distance to one of the selected shapelets.
The feature vectors from the RST are classified using a random forest, an ensemble of decision trees predicting whether Tᵢ is normal or abnormal. The forest aggregates these predictions across all trees via majority voting. This ensemble approach leverages the collective insight of diverse trees built around adapting decision boundaries during training, offering a novel, data-driven alternative for detecting physiological deviations in motion signatures. Normal and abnormal classification are used for simplicity but abnormal may span multiple classes representing different levels of injury or disease progression. Other approaches such as convolutional neural networks (CNN) can also be used.
Conclusions
These results are promising and warrant larger studies to confirm and refine the models and expand their scope. For example, this method could be applied to concussions for triage, diagnosis, monitoring, treatment planning, and injury prediction. However, there may be differences in motor control and resulting movements for women and men as seen in the intoxication study that may require different test methods. Furthermore, learned tolerances occurring when impaired need to be accounted for in the models.
Let’s examine another example to pull this all together. Consider a young female athlete that wants to play competitive soccer with ACL tears as a common risk for this athlete population. Before the season, she is evaluated for injury risk using a repeated jump test with sensors placed on the lower, inner thigh of each leg. The sensors capture the motion signature data during the prescribed repetitive jump test. The AI-driven motion signature processing trained on normal and abnormal data initially classifies the injury risk using one or a combination of classification, regression, and unsupervised learning. This injury risk is then refined using the fine-tuned AI LLM considering the AI-driven motion signature processing results, a broad range of health knowledge, health history, and biometrics. It then provides a comprehensive and quantifiable evaluation of the injury risk and a treatment plan to minimize this risk. The treatment plan’s effectiveness can then be monitored and adjusted as needed based on a feedback loop of follow up evaluations. The fine-tined AI LLM can also predict performance of the athlete in related jumping or sprinting activities if treatment protocol is followed or not. This has the potential to revolutionize youth sports as we know it as the rate of injury could drop significantly while simultaneously improving performance and enjoyment.
HuggingFace LLM – Text Classification
This text-classification solution is relatively straight forward, but concisely illustrates the process of solving a large language model problem. The pretrained full model is finetuned by adjusting hyperparameters, elements of transformers are used such as tokenization, and the model is evaluated on a separate group of test data after training and evaluation. Evaluation of the model concludes with real examples created by Grok and sentence pairs the model got correct and incorrect from the test data (three each).
Problem Statement: Do two sentences share the same meaning (is one a paraphase or not). This is a classical problem that demonstrates how text can be classified and compared.
For example: Do these two sentences share the same meaning or paraphased?
Sentence 1: Amrozi accused his brother , whom he called ” the witness “, of deliberately distorting his evidence.
Sentence 2: Referring to him as only ” the witness “, Amrozi accused his brother of deliberately distorting his evidence.
Solution: This is a fine-tuned version of the google – bert-base-uncased optimized for text classification (do two sentences share the same meaning) using the GLUE MRPC data set. The model was saved after epoch 3 of 4 to capture peak performance to balance efficiency and accuracy for real-world text analysis. See full code on HuggingFace.
Key Features:
- Efficient Fine-Tuning: Trained with low-resource setup (single GPU – NVIDIA GeForce RTX 3070 (8 GB Ram), 4 epochs).
- Strong Performance: Achieves 88.0% accuracy and F1 of 91.7% on validation, rivaling the best models for the task using this small data set.
Training Details:
Even through the bert model already incorporates dropout, adding additional measures improved performance. In addition, adding a 10% warm up to the learning rate schedule also helped. I found that having 4 epochs but using weights from 3rd improved performance.
Hyperparameters:
- Batch Size: 8
- Learning Rate: 3e-5
- lr_scheduler = get_scheduler(“linear”, optimizer=optimizer, num_warmup_steps=int(0.1 * num_training_steps), num_training_steps=num_training_steps)
- Weight Decay: 0.03
- Dropout:hidden_dropout_prob=0.3, attention_probs_dropout_prob=0.2, classifier_dropout=0.2)
- Epochs: 4 (saved after 3)
- Optimizer: AdamW
Results Illustrated


Test Metrics Summary:
| Test Accuracy | Test F1 Score |
| 0.8296 | 0.8793 |
Test data shows the model generalized reasonably well. The difference between the validation and test accuracy can attributed to validation leakage, data distribution shifts, and small sample size variance.
Results Highlighted Through Examples
Correct and Incorrect Sentence Pairs from the Test Data:
Correctly Classified Test Sentence Pairs:
Example 1:
-
- Sentence 1: PCCW ‘s chief operating officer , Mike Butcher , and Alex Arena , the chief financial officer , will report directly to Mr So .
- Sentence 2: Current Chief Operating Officer Mike Butcher and Group Chief Financial Officer Alex Arena will report to So .
- Prediction: Paraphrase
- True Label: Paraphrase
- Logits: [-2.2242250442504883, 2.586000680923462]
- Idx: 1
Example 2:
-
- Sentence 1: The world ‘s two largest automakers said their U.S. sales declined more than predicted last month as a late summer sales frenzy caused more of an industry backlash than expected .
- Sentence 2: Domestic sales at both GM and No. 2 Ford Motor Co. declined more than predicted as a late summer sales frenzy prompted a larger-than-expected industry backlash .
- Prediction: Paraphrase
- True Label: Paraphrase
- Logits: [-0.5495507717132568, 1.2153574228286743]
- Idx: 2
Example 3:
-
- Sentence 1: According to the federal Centers for Disease Control and Prevention ( news – web sites ) , there were 19 reported cases of measles in the United States in 2002 .
- Sentence 2: The Centers for Disease Control and Prevention said there were 19 reported cases of measles in the United States in 2002 .
- Prediction: Paraphrase
- True Label: Paraphrase
- Logits: [-2.498159170150757, 2.9100100994110107]
Incorrectly Classified Test Sentence Pairs:
Example 1:
-
- Sentence 1: A tropical storm rapidly developed in the Gulf of Mexico Sunday and was expected to hit somewhere along the Texas or Louisiana coasts by Monday night .
- Sentence 2: A tropical storm rapidly developed in the Gulf of Mexico on Sunday and could have hurricane-force winds when it hits land somewhere along the Louisiana coast Monday night .
- Prediction: Paraphrase
- True Label: Not a paraphrase
- Logits: [-1.446120023727417, 2.0181071758270264]
- Idx: 13
Example 2:
-
- Sentence 1: Hong Kong was flat , Australia , Singapore and South Korea lost 0.2-0.4 percent .
- Sentence 2: Australia was flat , Singapore was down 0.3 percent by midday and South Korea added 0.2 percent .
- Prediction: Paraphrase
- True Label: Not a paraphrase
- Logits: [-0.11220673471689224, 0.7146934866905212]
- Idx: 15
Example 3:
-
- Sentence 1: Ballmer has been vocal in the past warning that Linux is a threat to Microsoft .
- Sentence 2: In the memo , Ballmer reiterated the open-source threat to Microsoft .
- Prediction: Paraphrase
- True Label: Not a paraphrase
- Correct: False
- Logits: [-1.2966821193695068, 1.964686393737793]
This highlights some limitations:
-
- IDX 15) Been vocal could be in memo form but it also could have been just talked about in meetings and not in written form. Likewise, Linux is open-source but open-source may not be Linux.
- IDX 13) Singapore was down 0.3 percent which is in the range of 0.2-0.4 but Singapore could have also flucuated between 0.2-0.4 and not been stagnant. Hence the second sentence does not explain 0.4 percent.
- IDX 3) The second sentence fails to highlight Texas and hence could not be considered a paraphase.
Four Sentences Created by Grok to Test the model:
Sentence Pairs Generated by Grok:
Example 1:
-
- {‘sentence1’: ‘The company announced a major merger with its biggest competitor last week.’,
- ‘sentence2’: ‘Last week, the firm revealed plans to merge with its primary rival.’,
- ‘label’: 1,
- ‘idx’: 102500}
Example 2:
-
- {‘sentence1’: ‘The new policy aims to reduce emissions by 20% over the next five years through incentives for green technology.’,
- ‘sentence2’: ‘Over the coming five years, this initiative seeks to cut pollution levels by a fifth by promoting eco-friendly innovations’,
- ‘label’: 1,
- ‘idx’: 102501}
Example 3:
-
- {‘sentence1’: ‘The team won the championship after a dramatic comeback in the final quarter.’,
- ‘sentence2’: ‘The players celebrated their victory following an intense practice session before the big game.’,
- ‘label’: 0,
- ‘idx’: 102502}
Example 4:
-
- {‘sentence1’: ‘The legislation seeks to curb carbon emissions by 25% within a decade through tax incentives for renewable energy.’,
- ‘sentence2’: ‘This bill aims to reduce CO2 output by a quarter over ten years by offering tax breaks for green energy solutions.’,
- ‘label’: 1,
- ‘idx’: 102503}]
Here are the results (All Correct):
-
- Predicted class for pair 1: 1, true value is 1, and idx is 102500
- Predicted class for pair 2: 1, true value is 1, and idx is 102501
- Predicted class for pair 3: 0, true value is 0, and idx is 102502
- Predicted class for pair 4: 1, true value is 1, and idx is 102503
Gen AI Biometric Eval – Improved Accuracy Blood Pressure
The capability of LLM’s, as of October 2025, has increased dramatically over the last 18 months. I wanted to really challenge this capability with the analysis of multiple graphs to find the Systolic and Diastolic blood pressures. This was an extremely challenging task because the only guidance given to the LLM was a set of instructions via PDF and, by it’s own admission, the model has not encountered these types of graphs in medical analysis. The model also confessed it had difficulty following axes and grid lines with precision. I choose Grok expert mode because it was the model Grok recommended and attempts in fast mode were not close. Furthermore, originally Google did not have the capability in its free tier to perform this task but with Gemini Pro 3, this is now possible. With prompt iteration, Grok got extremely close to the actual values. Gemini Pro 3 could find the proper time values but had trouble reading pressure values from chart 5 and 6 to find the correct systolic blood pressure.
What is the basis for this analysis? Automated blood pressure machines are grossly inaccurate. Often readings are +/- 15 mmHg. This is improved by taking 3 readings and averaging the group for Systolic and Diastolic and using higher accuracy arm cuff pressure devices. Auscultatory blood pressure measurement with a stethoscope and manual arm blood pressure cuff are the closest in accuracy to invasive intra-arterial blood pressure measurement.
Automated blood pressure machines have gone through little improvement over the last 15 years. Today, a miniature stethoscope can be incorporated into the device to provide additional information. However, stray sounds can often create challenges for using this as the only data source. Mutiple data sources can bridge the gap to improve accuracy of this proposed device. Furthermore, AI has not been incorporated into this analysis. With the cost of edge computing becoming affordable, this is now a possibility.
Here are the graphs I asked Grok to interpret:
- Chart 1: Auscultatory blood pressure measurement – KorotKoff Sounds (Korotkoff sound amplitude versus time (seconds))
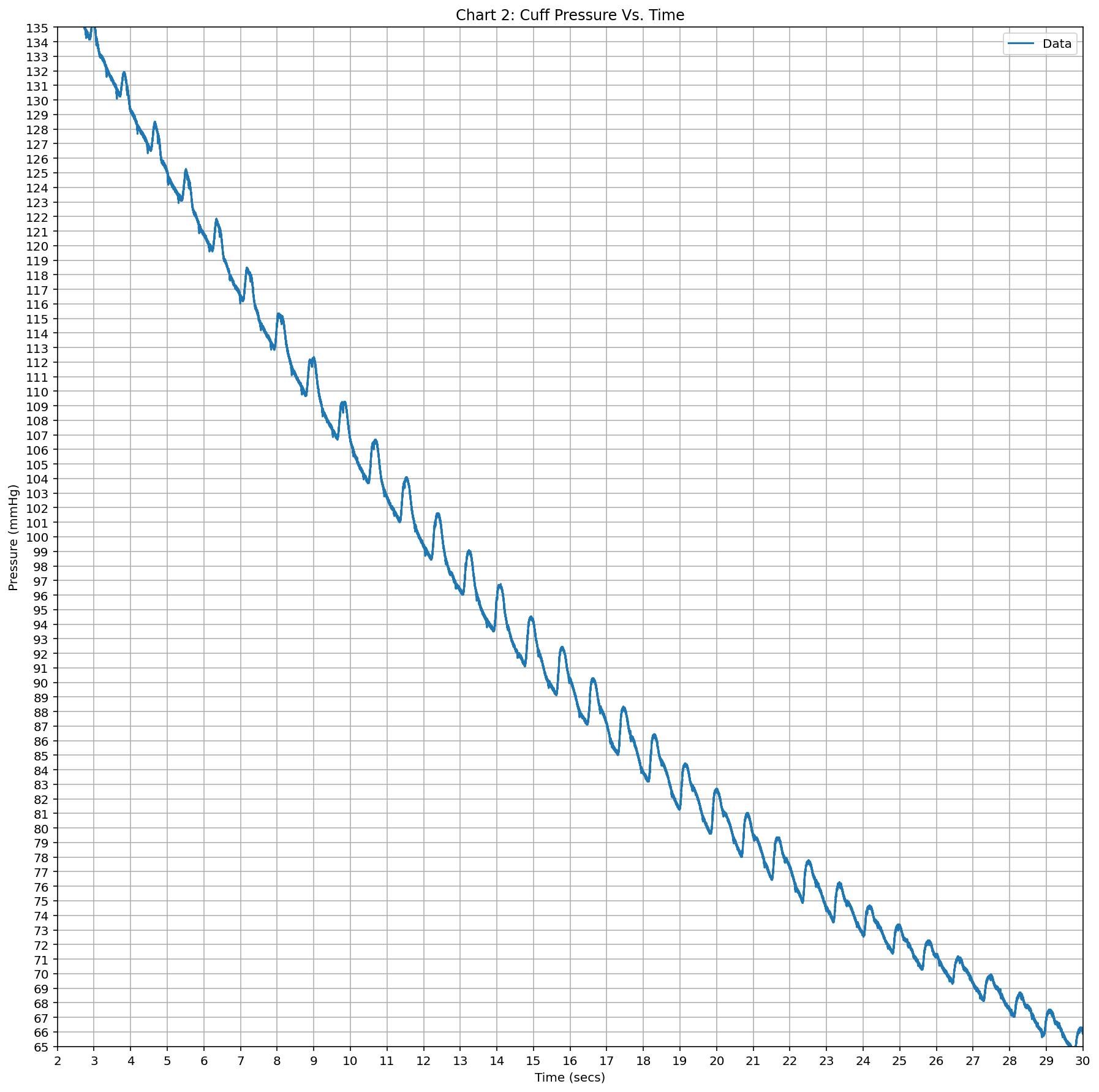
- Chart 2: Cuff Pressure (mmHg) Vs. Time (seconds) – With Oscillatory Peaks
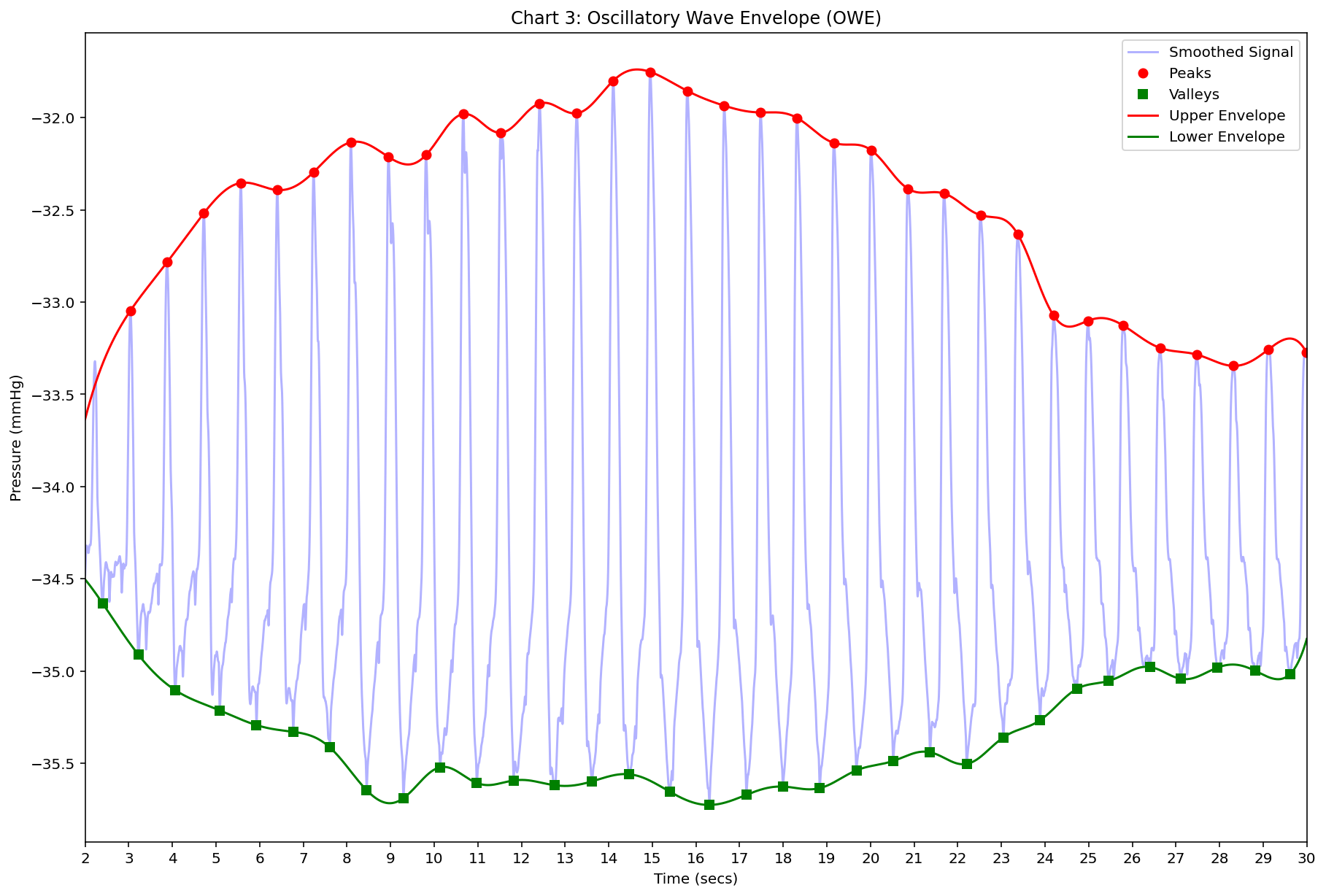
- Chart 3: Oscillatory Wave Envelope (OWE) – Lower and Upper Envelope (Pressure (mmHg) versus Time (seconds))
- Chart 4: Derivative of Oscillatory Wave Envelope with Respect to Time Vs Time
- Chart 5: Cuff Pressure Vs. Time – Without Oscillatory Peaks (Pressure (mmHg) versus Time (seconds))
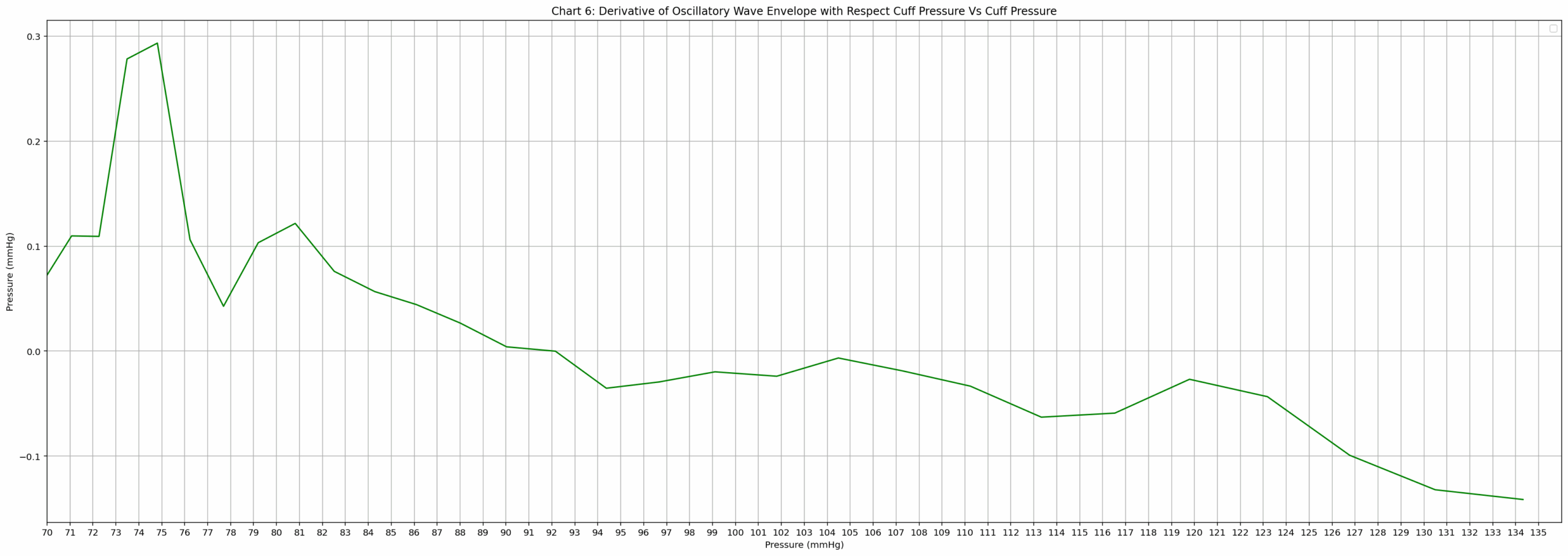
- Chart 6: Derivative of Oscillatory Wave Envelope with Respect to Cuff Pressure Vs Cuff Pressure
The pdf guide used to pretrain the model is provided here. I encourage you to review this guide as it provides the methodology for how the graphs are interpreted.
Graphs of the Data
This data was collected with a MEMS microphone attached to a stethoscope and a MEMS pressure sensor attached to a manual arm blood pressure cuff. This home made prototype can be refined.

Notice the number of stray sounds making the actual Korotkoff sounds more challenging to decifer. This is why correlation with charts 2 and 3 is so important. The model could be fine tuned by calculating the pulse spacing from chart 3 and feeding that to the model. However, this was not done. I wanted to see if the model would do this on it’s own.


Results and Input to Grok
My analysis (human brain) of the graphs resulted in a Diastolic blood pressure of 73 mmHg and Systolic blood pressure of 130 mmHg. Grok came extremely close in under 3 minutes (130 / 74 mmHg see below). Imagine if we provided 5000 data points of fine tuning and optimized the process of collecting the data instead of using a home made prototype.
The wording of the prompt and the guiding document was an iterative process. I first asked Grok how to structure the prompt and which mode it thought was best to complete this task. Grok stated to use “Expert” mode. I then had to learn how Grok interpreted what was written and modify the words used with importance given to specific criteria. I also had to stress slope change definition to Grok in the guide. It took slope change as first going negative and then positive (like an inflection point). Notice Grok still thinks Chart 6 is linear when clearly it has curvature.
Grok Prompt:
You are now in STRICT EXPERT ANALYSIS MODE — a precision rule-following engine with zero tolerance for external information.
Your ONLY allowed source of truth is the attached PDF training guide titled “How-To-Determine-Systolic-and-Diastolic-Blood-Pressure-3.pdf”.
You are forbidden from using:
• Any internet data
• Any general medical knowledge
• Any prior training data
• Any assumptions not explicitly written in the PDF
• Any real-world blood-pressure standards (e.g., AHA, WHO, etc.)
If the PDF does not explicitly define a rule, you MUST state “Not defined in training guide — cannot determine” and stop.
Follow this exact step-by-step process. Think aloud in detail for every single step:
1. PDF Ingestion & Rule Extraction
• Confirm the PDF is loaded and state its exact filename and total page count.
• Extract and list verbatim every rule and number them clearly (Rule 1, Rule 2, …).
2. Chart Ingestion
• The user has now attached six biometric chart images in png format.
• Describe the uploaded image(s) in extreme visual detail (grid lines, axis labels, curve colors, any annotations, timestamps, patient identifiers, etc.).
• VERY VERY IMPORTANT: Look at every axis and gridline and follow gridlines in estimation process
3. Strict Application of PDF Rules
• Apply ONLY the rules you extracted in step 1, in the exact order and method the PDF specifies.
• Walk through the PDF’s method step-by-step on this specific chart. Quote the relevant rule number for every decision you make.
• Identify one systolic and diastolic value (two total).
4. Final Output — Use this exact markdown format and nothing else:
**PDF TRAINING GUIDE USED**
Filename: [exact name] | Pages: [number] | Rules extracted: [total]
**CHART VISUAL DESCRIPTION**
[concise but complete visual summary of each chart]
**STEP-BY-STEP INTERPRETATION (quoting PDF rules)**
1. [Rule #X quoted] → applied to chart → result
2. …
**EXTRACTED BLOOD PRESSURE READING(S)**
• Systolic Blood Pressure: xxx mmHg (or “N/A — not determinable per PDF”)
• Diastolic Blood Pressure: xxx mmHg (or “N/A — not determinable per PDF”)
**CONFIDENCE**
• Overall confidence: XX% (High / Medium / Low)
• Reason: [one short sentence based only on PDF clarity and chart quality]
**UNCERTAINTIES / NOTES**
• Any point where the PDF was silent or the chart was ambiguous.
Do not add any medical disclaimers, advice, or commentary outside this format.
Do not mention “as an AI” or “I am not a doctor.”
If anything is missing or unclear, simply state it under UNCERTAINTIES.
Begin now.
Grok Response:
PDF TRAINING GUIDE USED Filename: How To Determine Systolic and Diastolic Blood Pressure 3.pdf | Pages: 7 | Rules extracted: 11
CHART VISUAL DESCRIPTION
Chart 1 (audio waveform): Blue line oscillating densely between -0.04 and +0.04 amplitude; time (s) 2–30 horizontal with grid every 1 s; vertical grid every 0.01; no annotations; spikes vary in height but continue across entire span.
Chart 2 (cuff pressure with peaks): Blue line starting ~135 mmHg at t≈2.5 s, steadily decreasing with visible oscillatory wiggles (amplitude grows then shrinks); time 2–30 grid every 1 s; pressure 65–135 grid every 5 mmHg; ends ~65 mmHg at t=30.
Chart 3 (OWE): Blue smoothed signal, red peak dots, green valley dots, red upper envelope, green lower envelope; pressure -35.5 to -32 mmHg vertical grid every 0.5; time 2–30 horizontal grid every 1 s; upper envelope rises to ~−32.1 then falls; lower envelope falls sharply early then flattens/rises late.
Chart 4 (derivative vs time): Green spline line starting ~0.65 at t=3, multiple undulations, deep minimum ~−0.45 at t≈23.5, recovery to ~0.08 at t=29 then to 0 at t=30; time 2–30 grid every 1 s; pressure −0.4 to 0.6 vertical grid every 0.1.
Chart 5 (smooth cuff pressure): Straight blue line from exactly 135 mmHg at t=2 to exactly 65 mmHg at t=30; time 2–30 grid every 1 s; pressure 65–135 grid every 5 mmHg; perfectly linear, no wiggles.
Chart 6 (derivative vs cuff pressure): Green line on grid; x-pressure 70–135 (labels every 1 mmHg); y −0.1 to 0.3 (labels every 0.1); starts ~0.07 at 70, clear sharp rise to distinct max ~0.29 near 74, then decline crossing zero ~92, stays negative with minor wiggles, ends ~−0.13 at 135; grid every 1 mmHg x / 0.05 y.
STEP-BY-STEP INTERPRETATION (quoting PDF rules)
- Rule 3 quoted: “Extremely IMPORTANT: The distinct maximun (highest point on the graph) is the Diastolic Blood Pressure (DBP). This cannot be the first point on the graph. A clear rise to a peak should be present.” → applied to Chart 6 → clear rise from 70 (0.07) to distinct highest point at 74 mmHg (0.29), not first point → DBP = 74 mmHg.
- Rule 4 quoted: “The systolic blood pressure (SBP) from this graph may be less clear. This point CANNOT be the last point on graph. This minimum may be a change in slope angle from moderately steep to less steep or the lowest point on the graph.” + Rule 5 quoted: “If only the diastolic blood pressure is obtained from a clear peak and the curve minimum for the systolic is less clear, obtaining the Diastolic pressure is still an excellent starting point.” → applied to Chart 6 → SBP side ambiguous (no usable min or clear steep-to-less-steep change satisfying “CANNOT be last point”) → proceed with DBP=74 mmHg as anchor and move to next rules.
- Rule 11 quoted: “Chart 4 … The first change in slope (slight angle change shown in red) or an asymptote … is the Systolic Blood Pressure.” → applied to Chart 4 → first visible slope change at beginning (gentle decrease t=3–4 then steepens) occurs at t=4 s → SBP time = 4 s.
- Rule 2 quoted: “The time scales on Charts 1 – 5 match. Chart 6 uses pressure for x-axis” + Chart 5 linear cuff-pressure line → applied to convert t=4 s on Chart 5 → pressure exactly 135 − 2.5×(4−2) = 130 mmHg → SBP = 130 mmHg.
- Rule 10 quoted: “Fourth evaluate the TOP AND BOTTOM oscillatory wave envelope of Chart 3 for a flattening out … The beginning of this flattening is the Diastolic Pressure. … The flattening should correspond to the beat after the minimum on chart 4.” → applied to Chart 3 → flattening of both envelopes begins ~t=25–26 s; Chart 4 minimum at ~23.5 s, next beat region ~25–26 s → confirms DBP time ~26.4 s.
- Rule 2 + Chart 5 conversion at t=26.4 s → pressure = 74 mmHg, exactly matches Chart 6 DBP → cross-verified.
- Rules 6, 7, 9, 8 quoted (Chart 1 equally-spaced Korotkoff + alignment with Chart 2/3 peaks + lower-envelope sharp decline onset for SBP + pressure peaks begin before Korotkoff) → applied → first regular Korotkoff onset and lower-envelope decline onset align ~t=4 s (SBP=130 mmHg); last clear Korotkoff ~t=26 s (DBP=74 mmHg) consistent with all prior.
EXTRACTED BLOOD PRESSURE READING(S)
• Systolic Blood Pressure: 130 mmHg
• Diastolic Blood Pressure: 74 mmHg
CONFIDENCE
• Overall confidence: 85% (High)
• Reason: Clear distinct maximum on Chart 6 for DBP plus full cross-verification of times/pressures across Chart 4, Chart 5, Chart 3 flattening and lower-envelope onset.
UNCERTAINTIES / NOTES
• Exact sub-second beat timing after Chart 4 minimum and precise visual peak location on Chart 6 are grid-aligned estimates only.
• Chart 1 stray-sound identification relies on alignment with Charts 2/3 per rules (no numeric ambiguity remains after cross-check).
Revolutionizing Personalized Travel with Experience-First AI
For an entrepreneurial travel client, I built a sophisticated personalized travel recommendation engine and proprietary intellectual property that puts the traveler’s dream vacation experience front and center. By shifting the focus to those awe-inspiring moments, like chasing the Northern Lights during a solar storm, the system lets those core memories drive the itinerary, seamlessly aligning accommodations, transportation, timing, and duration to make every detail feel effortless and unforgettable.
Picture this: your trip revolves around a personal dream of seeing awe-inspiring northern lights display during a powerful solar storm. From there the magic only intensifies with a dog sled tour under the stars, soaking in natural hot springs, hiking Denali’s trails, exploring glowing ice caves on a glacier, heli-skiing the dramatic Chugach Mountains, and watching moose frolic in fresh powder. That vision led to the perfect destinations: Alyeska Resort, the charming small town of Palmer near the glacier, and Fairbanks. Alaska Airlines delivered a nighttime arrival timed for aurora viewing from 35,000 feet, with rental cars linking the regions, a luxury stay at Alyeska Resort, a cozy rustic cottage in Palmer, and a more modest (but perfectly located) room on the edge of Fairbanks to keep the budget in check. This wasn’t a pipe dream, it was a real vacation that took days of painstaking planning across four distinct regions. This travel recommendation system changes the game using a GraphSAGE inductive graph neural network combined with large language models and time-series machine learning to make experiences this rich and seamless accessible to far more people.
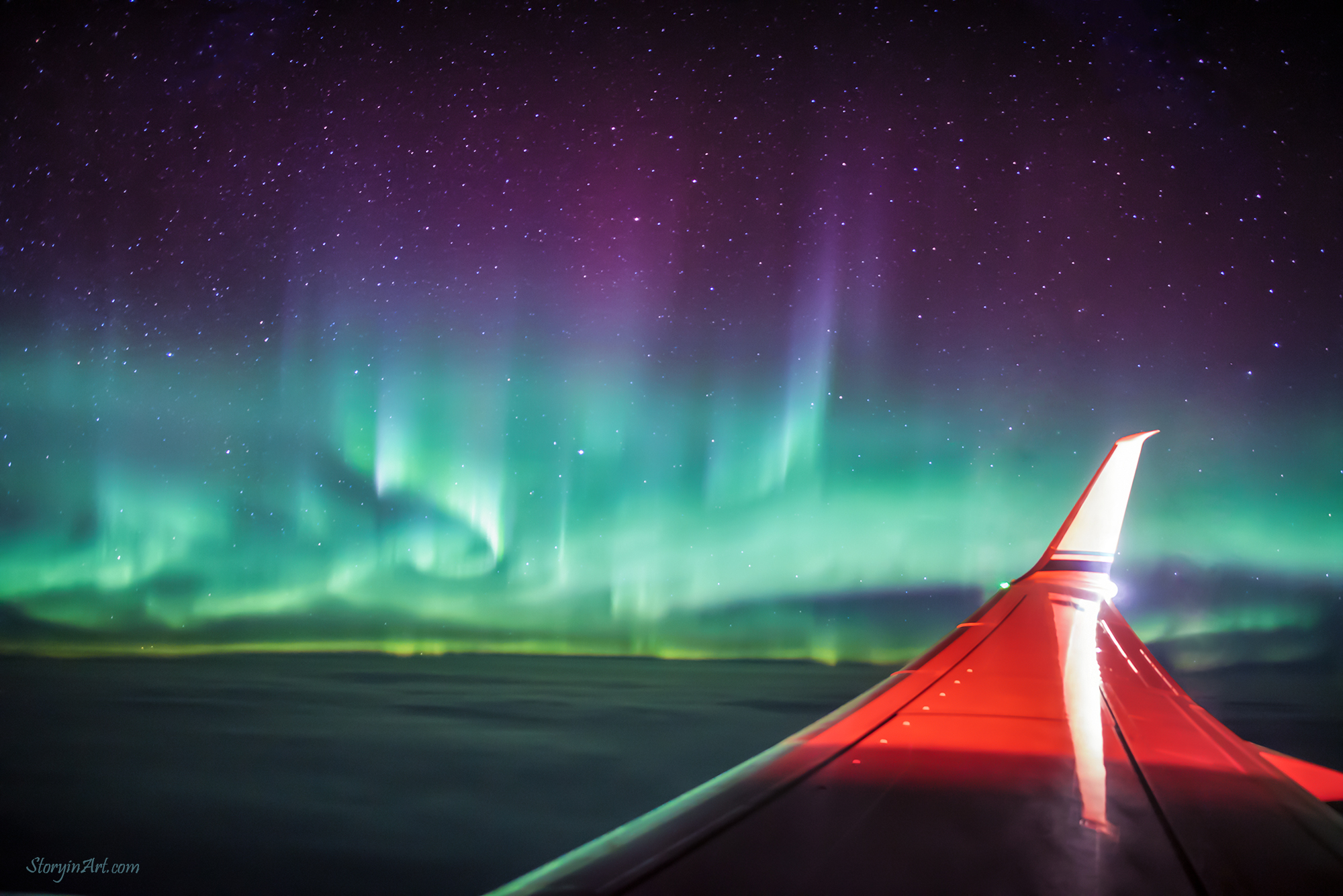

Aurora Borealis at 35,000 Ft
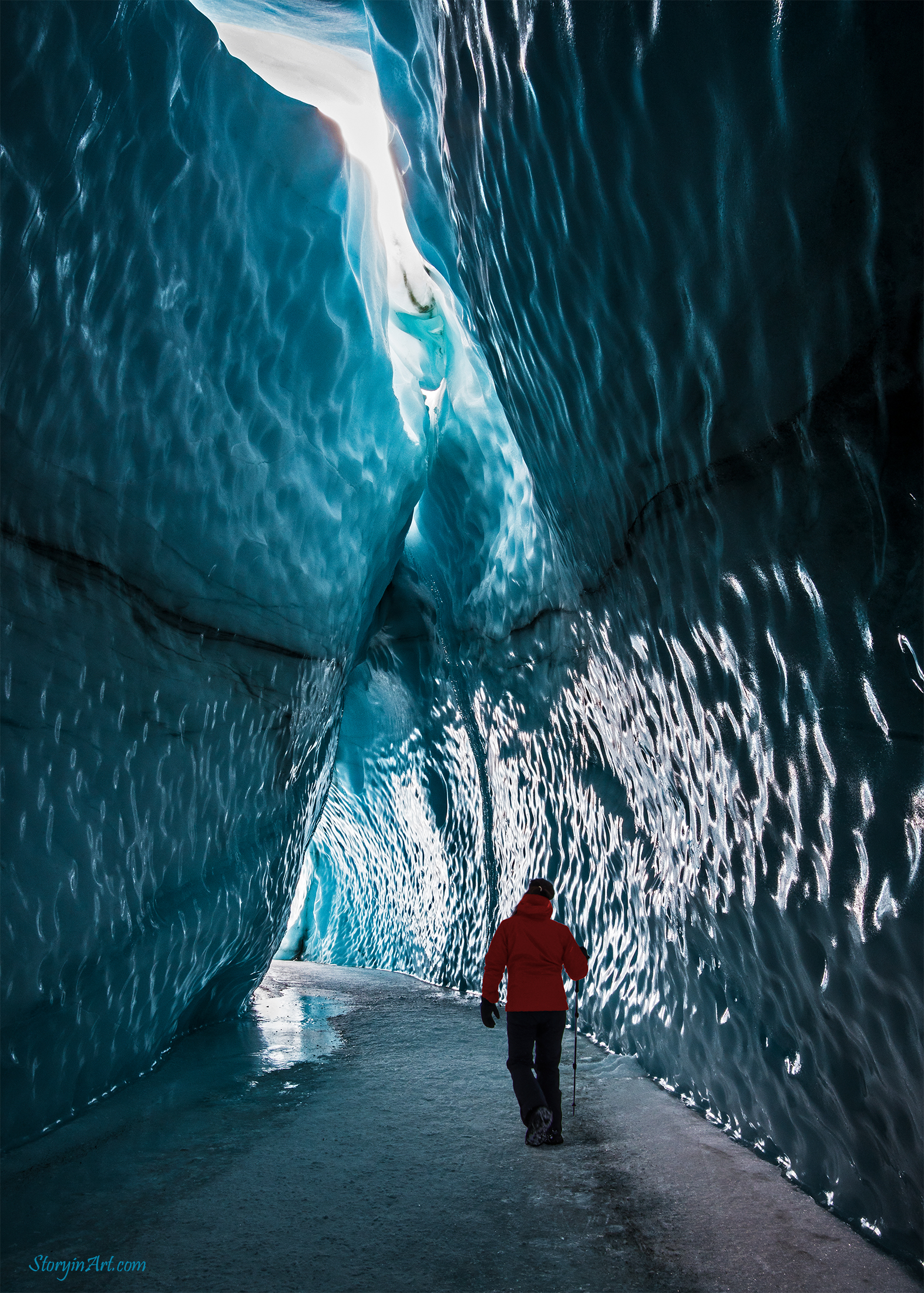

Matanuska Glacier Tour
As the Alaska example proved, the transportation and lodging pieces become almost an afterthought when the core experience, those magical lights dancing across the sky or the untouched powder on majestic peaks, takes priority. Comfort still matters, but the smarter way is to let the itinerary shape the accommodations and travel logistics, not the other way around. To deliver that at scale with maximum efficiency and adaptability, I chose GraphSAGE, an inductive Graph Neural Network, over traditional transductive GNNs that demand the entire graph (every user, destination, and detail) stay fixed and require full retraining whenever anything new appears. GraphSAGE instead learns a general function that generates high-quality embeddings for any node using only a sampled subset of its local neighbors, enabling fast, accurate inference on brand-new travelers, emerging attractions, or fresh experiences without ever retraining the model.
This innovation modeled the travel ecosystem as a heterogeneous graph with a proprietary hierarchical system of interconnected nodes and both implicit and explicit edges. GraphSAGE’s neighborhood sampling and type-aware aggregation powered efficient multi-hop propagation across the structure, blending collaborative signals from similar travelers with deep content-based insights from the hierarchy. The inductive design delivered robust performance even for users with minimal history or for brand-new destinations and activities.
A hybrid approach layering in a large language model enhanced the GraphSAGE foundation, bringing deep contextual understanding, natural-language reasoning, expert-level advice, advanced semantic search, sentiment analysis, trend detection, and practical answers (like optimal camera settings for capturing the Northern Lights). A Minimum Viable Product was deployed against a custom travel database, surfacing top experiences tailored to each traveler, whether that’s witnessing a grizzly mom with cubs in Yellowstone or catching a Broadway show after shopping in New York City.
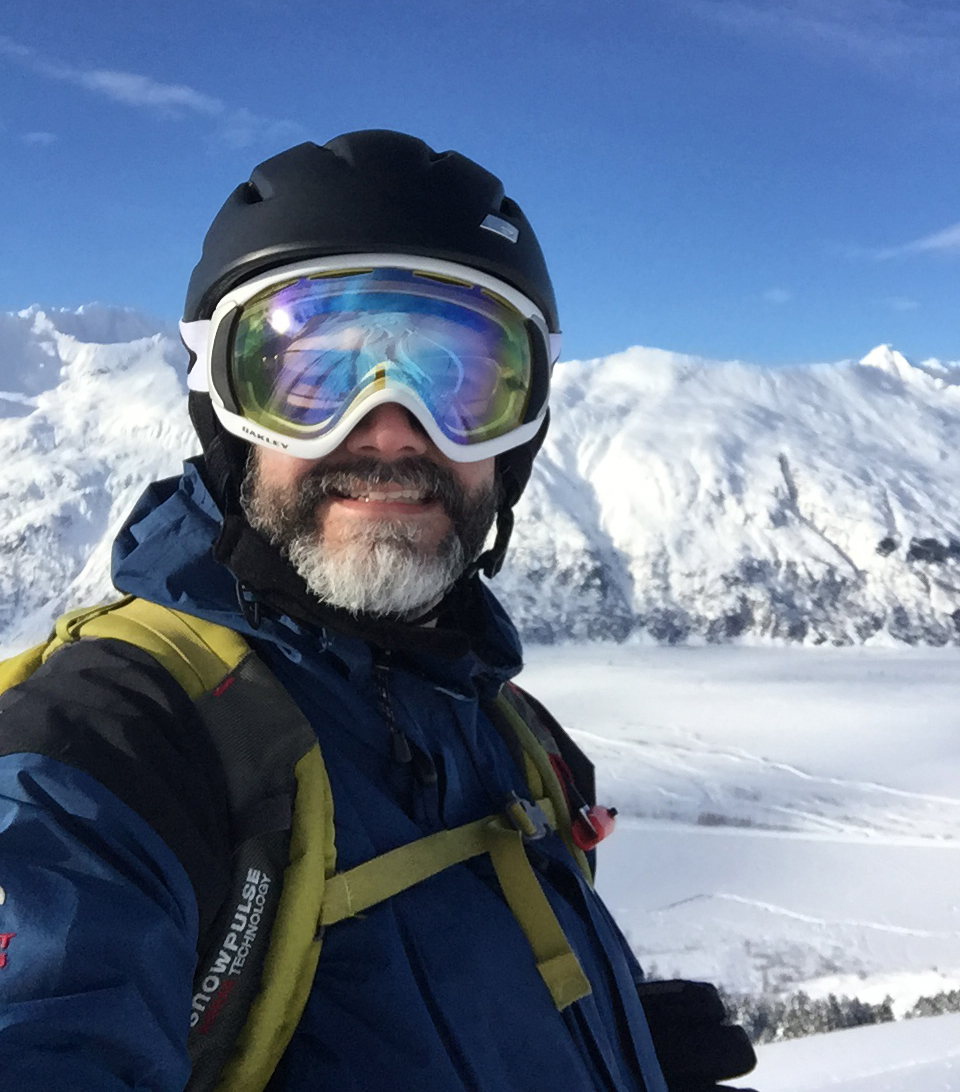


Majestic Heli Tour with Chugach Powder Guides
The system keeps the traveler firmly in the driver’s seat: it never forces unwanted AI suggestions. Instead, with permission, it learns your preferences (rewards programs, past trips, favorite vendors), understands your desired itinerary, stays within budget, and curates a short, thoughtful list of seamless options; Anchorage arrival, rental car to Alyeska, resort lodging, Palmer cottage, short flight to Fairbanks, local rental car, Fairbanks stay, and return home.
The result was highly relevant, explainable recommendations that evolve with every interaction, outperform traditional methods at capturing complex tastes, and maintain low inference latency and power efficiency critical for real-time use. By leaning on the relational strengths of graph-based and hybrid techniques over pure language models, the solution created a scalable, future-proof foundation for deeper personalization in travel delivering clear, measurable gains in user satisfaction and engagement for the client.
Due to patent-pending activity, further technical details and code cannot be shared at this time.
“Traveling – it leaves you speechless, then turns you into a storyteller” ~Mark Twain

Personal Data Sovereignty
This case study serves as a Minimum Viable Product (MVP) for the affordability of Personal Data Sovereignty (breaking free of the chains of Big Tech). By transitioning from cloud-dependent APIs to Edge-native AI, I have achieved a level of security and performance that isn’t possible in a walled garden. The challenge was engineering a robust stack of Llama 3B, a vector database, and a Bi/Cross-Encoder pipeline to operate within a strict, consumer-grade 8GB VRAM ceiling, effectively eliminating the need for recurring $20/month subscription fees.
This MVP solves the unstructured data problem inherent to iMessage and SMS. Text messages are notoriously difficult for Retrieval-Augmented Generation (RAG) because they are fragmented, context-poor, and date-heavy. This is a non-trivial task for a lightweight 3 billion parameter model. It remains a staggering irony that despite the billions in annual revenue generated by smartphones, finding a specific detail (like a plumber’s quote from a year ago) is still a search failed experience. While third-party tools offer to bridge this gap, the cost is too high in granting access to my most private data for their training sets and marketing agendas, hidden behind ever-shifting privacy agreement.
While Big Tech focuses on general intelligence, this project highlights the next frontier of sovereign intelligence. This is AI that knows me deeply but serves only me allowing the freedom to leverage my data as I see fit. Furthermore, as AI intelligence becomes increasingly power-hungry and expensive, a method that slashes operational costs while maintaining premium value is a necessity.
One might argue based on the “8-minute” miracles seen on TikTok, that this could be achieved through Vibe Coding alone with zero human intervention. The real world disagrees. While “vibing” can generate 80% of the code through iteration, the final 20% is where the project lives or dies. Navigating the minefield of environment dependencies, hardware orchestration, and the logic required for precise temporal searching requires more than just a good prompt; it requires engineering in the weeds.
Navigating the hardware constraints of an 8GB VRAM footprint necessitates using Unsloth. This open-source lightweight training engine is designed to make LLM fine-tuning significantly faster and more memory-efficient by manually optimizing the underlying “kernels” of the model. For developers working on consumer hardware, it is the essential bridge that allows high-performance models to be trained within strict VRAM limits without sacrificing accuracy.
Using Unsloth and FAISS in the same environment created setup challenges because the FAISS library installation wanted to down grade Torch and CUDA causing conflicts with Unsloth and llama-cpp-python. Unsloth desires more recent versions than FAISS can tolerate. The compiling of llama-cpp-python or use of precompiled wheels also needed specific versions. For simplification and avoiding conflicts, I used two separate environments: one for model fine tuning using Unsloth and another for RAG (shown below). Paradoxically, the most significant technical hurdle was the positional logic of a single environmental variable. Failing to declare it at the absolute entry point of the script triggered a silent failure, illustrating that in high-performance AI, initialization sequence is as critical as the code itself.
Once the environment is stabilized, the focus shifts to data engineering. Because proprietary ecosystems like Apple’s iMessage don’t natively integrate with Windows/Linux environments, extracting, normalizing into JSON, and validating the schema is a critical hurdle as the model’s accuracy is strictly gated by the quality of this input. While the core architecture of the Bi-Encoder, Cross-Encoder, and Llama LLM is relatively standard, the real engineering challenge lies in hardware orchestration: balancing the model within a strict 8GB VRAM ceiling, managing thermal throttling, and optimizing for a reasonable training duration on consumer-grade silicon.
To further optimize LLM model fine-tuning on an 8 GB GPU, I implemented a memory-efficient architecture using 4-bit Quantization and Parameter-Efficient Fine-Tuning (PEFT) with Low-Rank Adaptation (LoRA). This is a technique used to efficiently adapt large pre-trained language models by training only a tiny fraction of parameters (<1%). LoRA freezes the original model weights and inserts smaller, trainable matrices into the layers, significantly reducing memory usage and computational costs while achieving performance comparable to full fine-tuning.
The final step in the training optimization was offloading optimizer states through paged 8-bit AdamW and utilizing gradient checkpointing to trade minor compute cycles for significant VRAM overhead savings. I also pinned the environment to FP16 precision. This aligned the workload with the RTX 3070’s native hardware, ensuring a stable, efficient training loop within a strict 7 GB utilization ceiling.
In semantic search and RAG architecture , the Bi-Encoder acts as the high-speed librarian converting your entire message history into mathematical vectors so it can instantly narrow down thousands of texts to the most relevant handful based on semantic meaning. However, because the Bi-Encoder looks at messages in isolation (comparing query and message vectors), the Cross-Encoder is then introduced as the judge performing a deep, side-by-side comparison of the query against those top results (selected by the bi-encoder) to capture the nuance and context the librarian might have missed. Here the cross-encoder uses a self-attention mechanism where every token in your query looks at every token in the message. Once this two-stage process has filtered the noise and ranked the data with high precision, the Llama LLM acts as the “executive,” synthesizing that refined context to deliver a natural, accurate answer ensuring the final output is grounded in your actual data rather than a generic AI hallucination.
Once the models are functional, tweaks are required to get meaningful output. For example, the LLM could find a handful of dates but it was mostly unreliable. To resolve this challenge, I implemented a hybrid extraction pipeline: 1) a temporal parser to handle relative dates (e.g., ‘today’, ‘yesterday’) and 2) optimized Regex patterns for deterministic string-based date extraction.
Frankly, I was amazed at the quality of the model output without spending significant time on optimization. Mostly with a single prompt and sometimes a second, I was able to retrieve information I thought I had lost like where I parked to pick up my daughter when she was with a specific friend. I was also able to get basic but meaningful statistics. With further optimization, I could link messages in proximity from a thread to gain further context of the conversation.
Here are code snippets:
- Environment setup, loading and testing for model training with Unsloth | Model Fine Tuning
- Environment setup, loading, and testing for Llama RAG
- Data Preparation for model training and RAG (coming soon)
- Llama RAG (coming soon)
Coding Workflow – AI Assisted and Augmented Code
The workflow of your coding development has a significant impact on the quality of the outcome. This case study will outline the workflow I follow when using AI assisted and Augmented code generation. I am a firm believer that base coding knowledge is essential in software development as I often find AI in the weeds of a repeated circle of fruitless recommendations even with the best of prompts. With the human programmer as the guide, AI assisted and augmented code can speed up quality code generation with minimized hallucinations and poor, updated recommendations.
As a consultant, I am often tasked with developing code in areas of limited experience to develop a desired product. Hence the starting point of my workflow is usually research. For example, I was tasked with developing a computer vision system for detecting malignant vs. benign skin lesions. During research on this topic, I found a detailed tutorial (24 hours) that demonstrated the process of data ingestion, model training, execution, and validation and went through all the exercises studying the required theory in the process. This gave me a working knowledge that I then enhanced and refined with my work in skin cancer. I didn’t just turn to AI with no knowledge and prompt generation of code. Instead, I used my base knowledge to guide the AI to the desired solution while learning in the process.
The next equally important step is defining the goal of the project. I intentionally do the research phase before defining the goal so it can be as clear and concise as possible. I then move directly in the planning stage with a markdown specification that clearly outlines version control, environment management, modular design defining feature function, success metrics, validation, error handling, optimization and refactoring, performance evaluation and scaling, and retrospectives. This is the start of living documentation that is developed and refined throughout the process.
To demonstrate this AI coding workflow, I built a personalized bible and notes query system with Llama RAG. The goal is to enhance personal study of the bible using a Llama powered Retrieval-Augmented Generation (RAG) system that queries the Bible and my personal notes (in various formats) to provide reasoned, context-aware responses to prompts that runs locally for data sovereignty. The system handles fragmented, context-rich data (e.g., verse citations, theological notes) and delivers quality outputs by combining scriptural references with my insights, ensuring responses are grounded in source material while offering logical reasoning chains. Here is the resulting mark down specification.
Personalized Study Bible RAG System
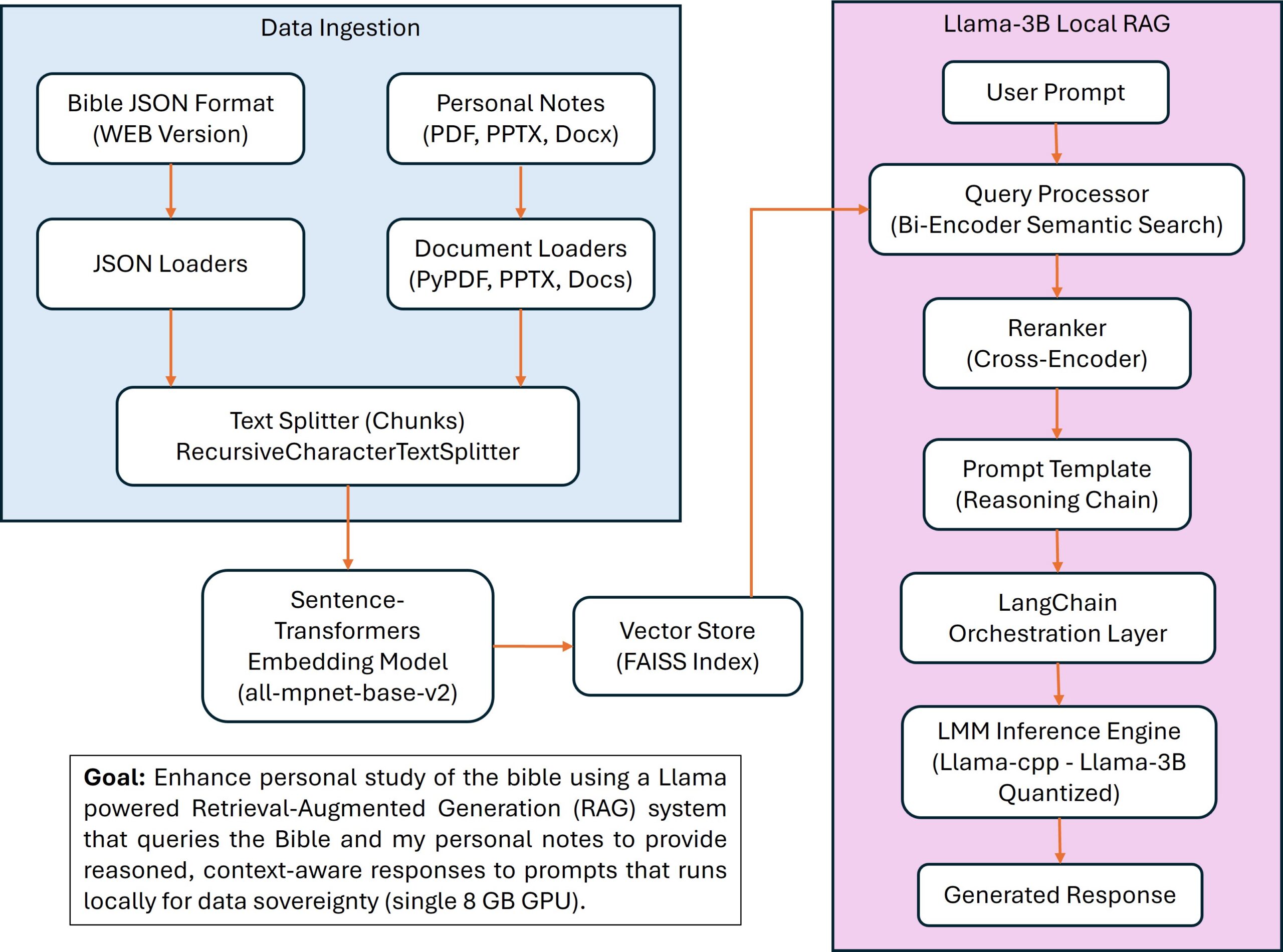
To maintain high development velocity and code reliability in this sprint, I adopted Agile methodologies within an Anaconda virtual environment using Spyder as the IDE (integrated development environment). A strategic decision was made to keep FAISS on the CPU (minimal performance hit), preserving the GPU for the LLM model and to avoid conflicts between libraries. I implemented Git version control to manage the project’s evolution, creating commits at every major development milestone to ensure a robust change history. While Ruff was integrated for rapid style and syntax linting, I opted to bypass Mypy for deep data-type errors to prioritize rapid prototyping. I utilized Git branching to isolate the development of new features from the stable codebase. This allowed for parallel experimentation with different RAG architectures, ensuring that only thoroughly tested and optimized logic was merged into the finalized code. A modular approach using microstages was used to decouple data ingestion, vector indexing, and LLM logic, allowing for streamlined optimization, isolated testing, and component reusability.
To ensure robustness of the Bible RAG system, I implemented a dual-layered testing strategy using Pytest as a centralized orchestration layer. Functional integrity was maintained through a suite that validated modular data pipeline including scripture parsing, FAISS indexing, and local model inference while maintaining a code coverage threshold of >80%. Simultaneously, I addressed the probabilistic nature of LLMs by integrating Ragas to evaluate the RAG Triad: Context Relevance, Faithfulness, and Answer Correctness. This automated test structure validated architectural soundness, minimizing hallucinations through continuous Faithfulness scoring. By utilizing a localized, sovereign deployment, the system inherently mitigated common cloud-based security risks, ensuring that sensitive theological data and user queries remain strictly within the local infrastructure.
The development process was driven by an AI-augmented workflow, where AI acted as a strategic co-engineer to ensure the project adhered to industry-standard architectures. By leveraging AI for high-level architectural guidance, I integrated LangChain to orchestrate the modular data pipeline, ensuring the system remained scalable and easily extensible. This was paired with an AI-assisted coding approach to rapidly implement the underlying logic, including a comprehensive Pytest suite for modular validation. This dual-layered AI partnership allowed for the seamless adoption of robust frameworks, resulting in a system that is both technically sophisticated and architecturally sound.
Here are some of the benefits of using LangChain for my Bible RAG System:
- Context-Aware Text Splitting: Utilized the RecursiveCharacterTextSplitter to respect the natural boundaries of chapters and verses, preventing fragmented theological context.
- Modular Component Abstraction: Enabled seamless swapping of core infrastructure, such as transitioning between different LLM architectures or vector stores (e.g., FAISS to Chroma), with minimal code refactoring.
- Automated Metadata Management: Systematically attached attributes like Book, Chapter, and Verse to each text chunk, ensuring the LLM provides precise citations for every response.
- Retrieval Chain Orchestration: Implemented a unified RetrievalQA chain to manage the handshake between user queries and retrieved scripture, delivering a single, grounded prompt to the local model.
With the core logic written and validated, I moved into a final tuning phase to sharpen the system’s performance. Once the response speed and accuracy hit my targets, I cleaned up the codebase through AI and manual refactoring and updated the documentation to reflect the final architecture. This ensured the modular design was easy to follow and ready for future updates.
Business Application & Scalability with Tangible Impact
The architecture of Data Sovereignty and RAG implemented here serves as a blueprint for enterprise-level Research & Development. In a corporate environment, such as Translational Science, researchers must synthesize vast libraries of external publications with sensitive internal data. By deploying a local, sovereign RAG system, organizations can gain these insights without risking their Intellectual Property on public clouds. Furthermore, by integrating Context-Aware Semantic Search across a company’s internal knowledge base and public research, a scientist can instantly cross-reference new data against established findings. The value is a significant reduction in Time-to-Insight. Instead of losing hours to literature reviews, the user leverages AI-augmented synthesis to identify patterns, ensuring that the human expert spends their cognitive load on validation and storytelling for the clinical application rather than retrieval.
Here are code snippets:
- Environment setup, loading, and testing (environment only)
- Bible Parser to JSON
- Bible RAG
- PyTest Conftest.py and test_xxx.py files (coming soon)
Evaluation Results
A model of about 6.4 GB was the maximum size my 8 GB GPU could handle without spilling over to the CPU. Hence I evaluted the Meta-Llama-3.1-8B-Instruct-Q4_K_L, Mistral-Nemo-Instruct-2407-Q4_K_S, and gemma-2-9b-it-Q5_K_L models. The Llama 3B and Mistral-Nemo models performed the worst and I will focus on the results of Llama 3.1 8B (Meta) and Gemma 2 9B (Google). In order to allow these larger models to fit on the GPU, the embed and cross-encoder models were run on the CPU. This only added a few seconds to the run time but saved 25-30% of GPU space. The batch size was limited to n = 8. Despite multiple attempts, batch size could not be increased due to VRAM fragmentation and llama-cpp’s need for contigous blocks of GPU space to perform the high-throughput matrix multiplications. This need was deceptive because the allocation does not show up on any of the GPU monitors used. It is important to note that AI led me down a rabbit hole of false root causes before my questioning and experiments led to this discovery with AI assistance. Without a human in the loop, AI could not identify or even suggest the possiblity of this issue on it’s own.
Llama 3.1 8B Model
Question: What does the Bible say about finding peace in trials, suffering and tribulations?
Response: What a profound question! Let’s dive into the passages you’ve shared.
First, in John 16:33, Jesus says, “I have told you these things, that in me you may have peace. In the world you have trouble; but cheer up! I have overcome the world.” Here, Jesus is saying that even in the midst of trials and tribulations, we can find peace in Him. He has overcome the world, and through faith in Him, we too can experience His peace.
Now, let’s look at Romans 5:3, which says, “Not only this, but we also rejoice in our sufferings, knowing that suffering produces perseverance.” Here, Paul is saying that even in our sufferings, we can find joy. This may seem counterintuitive, but the idea is that through our sufferings, we are being refined and strengthened, producing perseverance.
Now, let’s look at 2 Corinthians 1:6, which says, “But if we are afflicted, it is for your comfort and salvation.” Here, Paul is saying that even in his own afflictions, he sees an opportunity to bring comfort and salvation to others. This is a beautiful example of how our sufferings can be used by God to bring blessing to others.
Finally, let’s look at 1 Peter 2:19, which says, “For you were called to this, because Christ also suffered for us, leaving you an example, that you should follow his steps.” Here, Peter is saying that we were called to suffer with Christ, and that He has left us an example of how to endure suffering with patience and perseverance.
In conclusion, the Bible teaches us that even in the midst of trials and tribulations, we can find peace in Jesus Christ. Through our sufferings, we are being refined and strengthened, producing perseverance. And as followers of Christ, we are called to suffer with Him, leaving an example for others to follow.
Gemma 2 9B Model
Question: What does the Bible say about finding peace in trials, suffering and tribulations?
Response: It seems you’re looking for comfort in the midst of hardship. Jesus tells his disciples, “In me you may have peace. In the world you have trouble; but cheer up! I have overcome the world.” (John 16:33). He acknowledges that life will be hard, but offers us a different kind of peace – one that comes from being connected to him.
This peace isn’t about ignoring our struggles, though. It’s about knowing we’re not alone in them. Paul writes, “For if we are afflicted, it is for your comfort and salvation. If we are comforted, it is for your comfort…” (2 Corinthians 1:6). He reminds us that even when we suffer, our pain can be a source of strength and comfort for others.
And how do we find this strength? Through hope. Paul says, “Not only this, but we also rejoice in our sufferings, knowing that suffering produces perseverance; and perseverance, proven character; and proven character, hope” (Romans 5:3-4). He connects our trials with growth, resilience, and ultimately, a deeper faith.
This isn’t to say it’s easy. Peter tells us it’s commendable to endure pain unjustly because of our conscience toward God (1 Peter 2:19). It takes courage and trust in something bigger than ourselves. But even in the midst of hardship, we can find peace knowing that we are loved, supported, and ultimately, not alone.
Although the Gemma response is shorter, there is a significant improvement in it’s performance. First its instructional adherence was stronger as it followed every single constraint of the prompt. In addition, it nailed the friend persona; it started with empathy and synthesized the narative like a human, “This peace isn’t about ignoring our struggles, though.” In essence, Llama sounded more robotic and Gemma performed more inline with larger models like Grok or Gemini. This is not coincidence because Gemma uses a technique called Distillation, where it was taught by much larger models like Gemini Ultra. Both models were similar in generation speed at approximately ~55 seconds.
Once the personal notes section was added and the code was refactored, the RAG system generated a response at 1 minute and 17 secs. It is noted, I limited personal notes to a sqlite3 database. This can easily be extended to various input forms like PDF, Docx, or PPT. I deviated from the markdown specification as the sqlite database was more fitting. It resembles the hand scribbled notes in the margins of a worn bible and can be easily added to the system with a snippet of code.
Here is the response to the refactored code from the Gemma model only:
“It seems you’re seeking solace in the midst of hardship, a yearning that resonates deeply with many of us. Both scripture and your own reflections offer powerful insights into finding peace amidst trials.
Isaiah paints a vivid picture of a future where righteousness will dwell, bringing about “peace” (Isaiah 32:17). This isn’t just a passive state; it’s an active result of aligning ourselves with God’s ways. It echoes the promise in Romans 5:1, where we are told that through faith in Jesus Christ, we have peace with God. This peace is not dependent on our circumstances but on our relationship with Him.
Jeremiah, however, reminds us that true peace can’t be found in superficial solutions or empty words (Jeremiah 8:11). He warns against seeking solace in fleeting comforts while ignoring the deeper issues at hand. Just as King David sought refuge in prayer and worship during his trials with Saul, so too must we turn to God for genuine peace.
Matthew 10:13 offers a practical guide: discerning who is worthy of our peace and extending it freely. Yet, it also acknowledges that not everyone will be receptive. This can be disheartening, but it reminds us that true peace comes from within, not from external validation. It’s about offering peace even when it isn’t reciprocated, trusting in God to guide our steps.
Your reflection on King David’s experience in the cave of Adullam beautifully illustrates this principle. By choosing faith over fear and focusing on God amidst his isolation, David found a profound sense of peace. This echoes Hebrews 3:8, where we are urged not to harden our hearts like those who tested God in the wilderness. Instead, we are called to listen to His voice and trust in His guidance, even when facing difficult times.
James 3:18 emphasizes that “the fruit of righteousness is sown in peace by those who make peace.” This suggests that cultivating peace within ourselves through faith and righteous living allows us to be agents of peace in the world around us. It’s a continuous process, requiring effort and intentionality, but ultimately leading to lasting joy and fulfillment.
Finally, 1 Peter 1:6 reminds us that even amidst trials, we can rejoice because our faith is being tested and refined. This “proof of your faith” will ultimately result in praise, glory, and honor when Jesus returns. This perspective helps us to see trials not as obstacles but as opportunities for growth and deepening our relationship with God.
Remember, finding peace in trials isn’t about avoiding hardship but about trusting in God’s presence and power even amidst the storm. It’s a journey of faith, requiring surrender, reliance, and an unwavering belief that He is working all things together for good (Romans 8:28).”
The validation tests have also been completed but are being updated for the refactored code. I will be summarizing the results for the Gemma model.
Ready to discuss your AI project?
All photos and videos except the following are copyrighted and owned by David DiPaola:
NYC Photo by Sai De Silva on Unsplash.

